| functional annotation |
| Function |
B-box type zinc finger family protein |


|
| GO BP |
|
GO:0010100 [list] [network] negative regulation of photomorphogenesis
|
(13 genes)
|
IDA
|
 |
|
GO:0009640 [list] [network] photomorphogenesis
|
(71 genes)
|
IMP
|
 |
|
GO:0007623 [list] [network] circadian rhythm
|
(111 genes)
|
IEP
|
 |
|
GO:0010228 [list] [network] vegetative to reproductive phase transition of meristem
|
(173 genes)
|
IMP
|
 |
|
GO:0006355 [list] [network] regulation of transcription, DNA-templated
|
(1984 genes)
|
TAS
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0008270 [list] [network] zinc ion binding
|
(554 genes)
|
IEA
|
 |
|
GO:0003700 [list] [network] DNA-binding transcription factor activity
|
(1599 genes)
|
ISS
|
 |
|
GO:0005515 [list] [network] protein binding
|
(4605 genes)
|
IPI
|
 |
|
| KEGG |
|
|
| Protein |
NP_001031811.1
NP_001190958.1
NP_001320162.1
NP_001320163.1
NP_001328010.1
NP_195607.2
|
| BLAST |
NP_001031811.1
NP_001190958.1
NP_001320162.1
NP_001320163.1
NP_001328010.1
NP_195607.2
|
| Orthologous |
[Ortholog page]
BBX18 (ath)
LOC4347644 (osa)
LOC7463992 (ppo)
LOC7470030 (ppo)
LOC7480165 (ppo)
LOC11420536 (mtr)
LOC100193329 (zma)
LOC100257242 (vvi)
LOC100527369 (gma)
LOC100780544 (gma)
LOC100795117 (gma)
LOC100818604 (gma)
LOC100852825 (vvi)
LOC101264380 (sly)
LOC103835153 (bra)
LOC103842538 (bra)
|
Subcellular
localization
wolf |
|
nucl 4,
mito 3,
chlo 1,
cyto 1,
cyto_mito 1,
mito_plas 1
|
(predict for NP_001031811.1)
|
|
cyto 8,
nucl 1,
plas 1,
cysk 1,
cysk_nucl 1,
nucl_plas 1,
cysk_plas 1
|
(predict for NP_001190958.1)
|
|
cyto 9,
nucl 1,
plas 1,
nucl_plas 1
|
(predict for NP_001320162.1)
|
|
nucl 4,
mito 3,
chlo 1,
cyto 1,
cyto_mito 1,
mito_plas 1
|
(predict for NP_001320163.1)
|
|
nucl 6,
cyto_nucl 4,
mito 2,
chlo 1
|
(predict for NP_001328010.1)
|
|
cyto 9,
nucl 1,
plas 1,
nucl_plas 1
|
(predict for NP_195607.2)
|
|
Subcellular
localization
TargetP |
|
other 5
|
(predict for NP_001031811.1)
|
|
scret 7
|
(predict for NP_001190958.1)
|
|
scret 7,
other 4
|
(predict for NP_001320162.1)
|
|
other 5
|
(predict for NP_001320163.1)
|
|
other 5
|
(predict for NP_001328010.1)
|
|
scret 7,
other 4
|
(predict for NP_195607.2)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath04712 |
Circadian rhythm - plant |
2 |
 |
Genes directly connected with BBX19 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 12.2 |
LHY |
Homeodomain-like superfamily protein |
[detail] |
839341 |
| 12.1 |
CCA1 |
circadian clock associated 1 |
[detail] |
819296 |
| 10.6 |
RVE8 |
Homeodomain-like superfamily protein |
[detail] |
820117 |
| 10.4 |
BBX18 |
B-box zinc finger family protein |
[detail] |
816671 |
| 9.3 |
NDT1 |
NAD+ transporter 1 |
[detail] |
819362 |
| 7.7 |
NF-YA4 |
nuclear factor Y, subunit A4 |
[detail] |
818037 |
| 7.2 |
AT5G15950 |
Adenosylmethionine decarboxylase family protein |
[detail] |
831452 |
| 7.0 |
AT5G67140 |
F-box/RNI-like superfamily protein |
[detail] |
836849 |
| 5.7 |
AT1G19490 |
Basic-leucine zipper (bZIP) transcription factor family protein |
[detail] |
838535 |
| 3.8 |
AT1G52565 |
cytochrome P450 family protein |
[detail] |
841688 |
|
Coexpressed
gene list |
[Coexpressed gene list for BBX19]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
252917_at
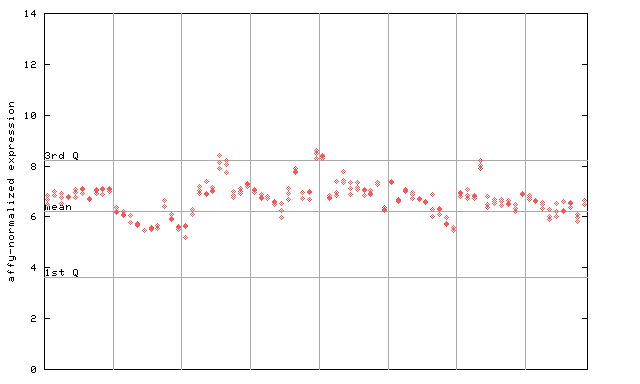
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
252917_at
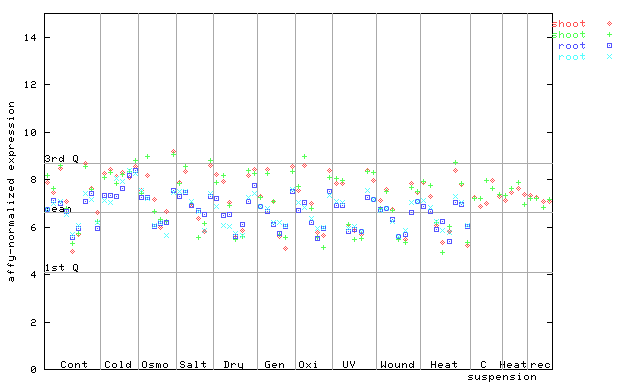
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
252917_at
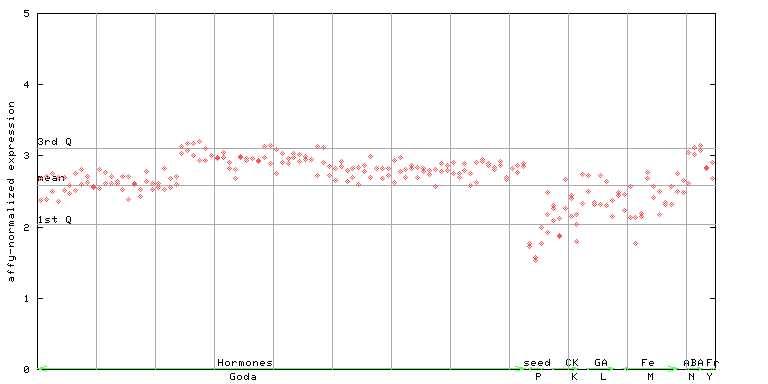
X axis is samples (xls file), and Y axis is log-expression.
|















