| functional annotation |
| Function |
3'-phosphoinositide-dependent protein kinase 1 |


|
| GO BP |
|
GO:0045860 [list] [network] positive regulation of protein kinase activity
|
(59 genes)
|
IDA
|
 |
|
GO:0046777 [list] [network] protein autophosphorylation
|
(173 genes)
|
IDA
|
 |
|
| GO CC |
|
GO:0005886 [list] [network] plasma membrane
|
(3771 genes)
|
ISM
|
 |
|
GO:0016020 [list] [network] membrane
|
(7839 genes)
|
IEA
|
 |
|
GO:0005634 [list] [network] nucleus
|
(10793 genes)
|
ISM
|
 |
|
GO:0005737 [list] [network] cytoplasm
|
(14855 genes)
|
IEA
|
 |
|
| GO MF |
|
GO:0004676 [list] [network] 3-phosphoinositide-dependent protein kinase activity
|
(1 genes)
|
IGI
|
 |
|
GO:0070300 [list] [network] phosphatidic acid binding
|
(13 genes)
|
IDA
|
 |
|
GO:0035091 [list] [network] phosphatidylinositol binding
|
(78 genes)
|
IDA
|
 |
|
GO:0004674 [list] [network] protein serine/threonine kinase activity
|
(801 genes)
|
IDA
|
 |
|
GO:0004672 [list] [network] protein kinase activity
|
(1017 genes)
|
IDA
|
 |
|
GO:0016301 [list] [network] kinase activity
|
(1362 genes)
|
ISS
|
 |
|
GO:0005524 [list] [network] ATP binding
|
(2003 genes)
|
IEA
|
 |
|
GO:0005515 [list] [network] protein binding
|
(4605 genes)
|
IPI
|
 |
|
| KEGG |
|
|
| Protein |
NP_001330324.1
NP_568138.1
NP_974730.1
|
| BLAST |
NP_001330324.1
NP_568138.1
NP_974730.1
|
| Orthologous |
[Ortholog page]
Pdk1 (sly)
AT3G10540 (ath)
LOC4324953 (osa)
LOC7468462 (ppo)
LOC25484865 (mtr)
LOC100264394 (vvi)
LOC100788780 (gma)
LOC100801891 (gma)
LOC101262698 (sly)
LOC103650947 (zma)
LOC103847282 (bra)
LOC103847461 (bra)
LOC103870456 (bra)
|
Subcellular
localization
wolf |
|
cyto 6,
chlo 1,
cysk 1,
nucl 1,
mito 1,
cysk_plas 1
|
(predict for NP_001330324.1)
|
|
cyto 6,
nucl 4
|
(predict for NP_568138.1)
|
|
nucl 7,
cyto 1,
extr 1,
pero 1,
cyto_E.R. 1,
cyto_plas 1
|
(predict for NP_974730.1)
|
|
Subcellular
localization
TargetP |
|
other 6
|
(predict for NP_001330324.1)
|
|
other 6
|
(predict for NP_568138.1)
|
|
other 6
|
(predict for NP_974730.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
Genes directly connected with PDK1 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 6.7 |
AT1G48230 |
nodulin MtN21 /EamA-like transporter family protein |
[detail] |
841243 |
| 6.2 |
AtPOT1a |
Nucleic acid-binding, OB-fold-like protein |
[detail] |
815069 |
| 5.9 |
AT5G50970 |
transducin family protein / WD-40 repeat family protein |
[detail] |
835170 |
| 5.7 |
AT5G50180 |
Protein kinase superfamily protein |
[detail] |
835083 |
| 5.7 |
AT5G57270 |
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein |
[detail] |
835832 |
| 5.5 |
ZW18 |
Putative serine esterase family protein |
[detail] |
842204 |
| 5.0 |
AT1G19690 |
NAD(P)-binding Rossmann-fold superfamily protein |
[detail] |
838556 |
| 4.8 |
AT1G28090 |
Polynucleotide adenylyltransferase family protein |
[detail] |
839702 |
| 4.8 |
MAN2 |
Glycosyl hydrolase superfamily protein |
[detail] |
816596 |
| 4.6 |
CLT1 |
CRT (chloroquine-resistance transporter)-like transporter 1 |
[detail] |
832058 |
|
Coexpressed
gene list |
[Coexpressed gene list for PDK1]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
250848_at
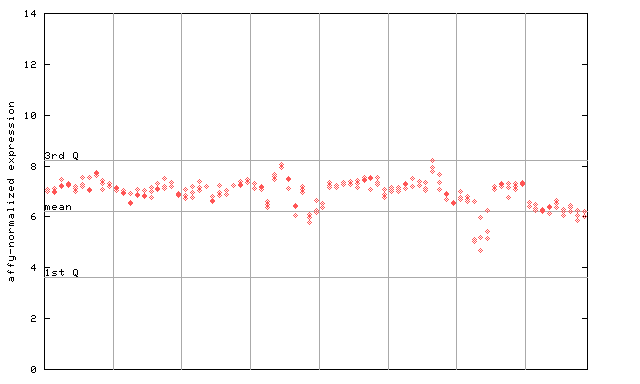
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
250848_at
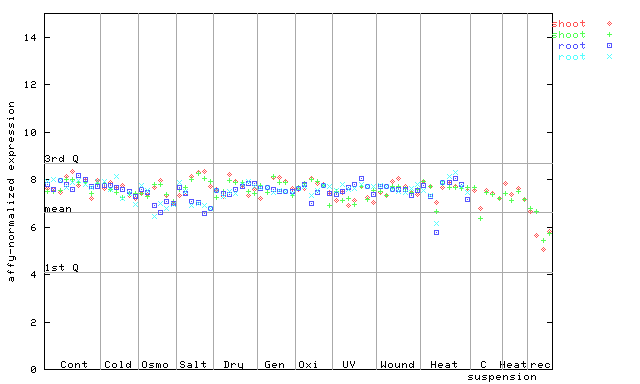
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
250848_at
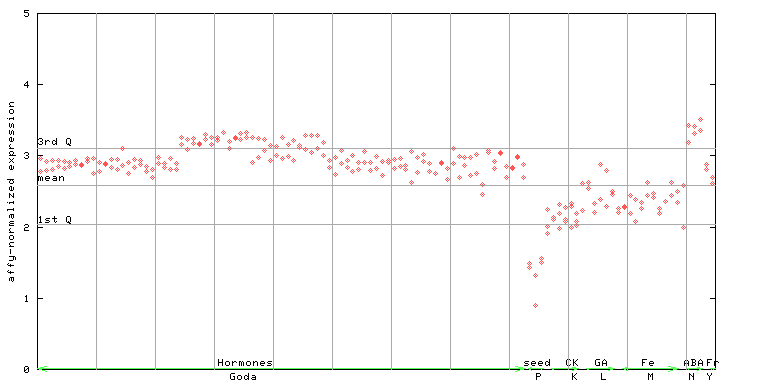
X axis is samples (xls file), and Y axis is log-expression.
|











