| functional annotation |
| Function |
DNA/RNA helicase protein |


|
| GO BP |
|
GO:0045003 [list] [network] double-strand break repair via synthesis-dependent strand annealing
|
(7 genes)
|
IDA
|
 |
|
GO:0036297 [list] [network] interstrand cross-link repair
|
(14 genes)
|
IGI
|
 |
|
GO:0009294 [list] [network] DNA mediated transformation
|
(26 genes)
|
IMP
|
 |
|
GO:0000724 [list] [network] double-strand break repair via homologous recombination
|
(68 genes)
|
IDA
IMP
|
 |
|
GO:0006325 [list] [network] chromatin organization
|
(338 genes)
|
IEA
|
 |
|
| GO CC |
|
GO:0005634 [list] [network] nucleus
|
(10793 genes)
|
IEA
ISM
|
 |
|
| GO MF |
|
GO:0004386 [list] [network] helicase activity
|
(157 genes)
|
IEA
|
 |
|
GO:0008270 [list] [network] zinc ion binding
|
(554 genes)
|
IEA
|
 |
|
GO:0005524 [list] [network] ATP binding
|
(2003 genes)
|
IEA
|
 |
|
GO:0003676 [list] [network] nucleic acid binding
|
(4187 genes)
|
IEA
|
 |
|
| KEGG |
|
|
| Protein |
NP_197667.1
|
| BLAST |
NP_197667.1
|
| Orthologous |
[Ortholog page]
LOC4329524 (osa)
LOC25487494 (mtr)
LOC100246473 (vvi)
LOC100794452 (gma)
LOC100794558 (gma)
LOC101267090 (sly)
LOC103653576 (zma)
LOC103856528 (bra)
|
Subcellular
localization
wolf |
|
nucl 5,
cyto 3,
chlo 1,
cyto_pero 1,
cyto_E.R. 1,
cyto_plas 1
|
(predict for NP_197667.1)
|
|
Subcellular
localization
TargetP |
|
other 7
|
(predict for NP_197667.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath03030 |
DNA replication |
4 |
 |
| ath03440 |
Homologous recombination |
3 |
 |
| ath03430 |
Mismatch repair |
2 |
 |
Genes directly connected with RAD5 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 9.1 |
TSK |
tetratricopeptide repeat (TPR)-containing protein |
[detail] |
821404 |
| 8.9 |
MCM6 |
minichromosome maintenance (MCM2/3/5) family protein |
[detail] |
834492 |
| 8.9 |
MEI1 |
transcription coactivator |
[detail] |
844068 |
| 7.5 |
MCM8 |
minichromosome maintenance 8 |
[detail] |
820123 |
| 7.0 |
MSH3 |
homolog of DNA mismatch repair protein MSH3 |
[detail] |
828659 |
| 6.7 |
MSH6 |
MUTS homolog 6 |
[detail] |
828147 |
| 6.3 |
AT5G07400 |
forkhead-associated domain-containing protein / FHA domain-containing protein |
[detail] |
830631 |
| 5.7 |
AT3G17340 |
ARM repeat superfamily protein |
[detail] |
820997 |
| 5.4 |
AT3G53680 |
Acyl-CoA N-acyltransferase with RING/FYVE/PHD-type zinc finger domain-containing protein |
[detail] |
824535 |
| 5.0 |
AT1G77620 |
P-loop containing nucleoside triphosphate hydrolases superfamily protein |
[detail] |
844097 |
| 5.0 |
Pol{lambda} |
DNA polymerase lambda (POLL) |
[detail] |
837592 |
| 4.8 |
AT5G08110 |
UBQ, helicase-c and DEAD-like helicase domain-containing protein |
[detail] |
830705 |
|
Coexpressed
gene list |
[Coexpressed gene list for RAD5]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
249907_at
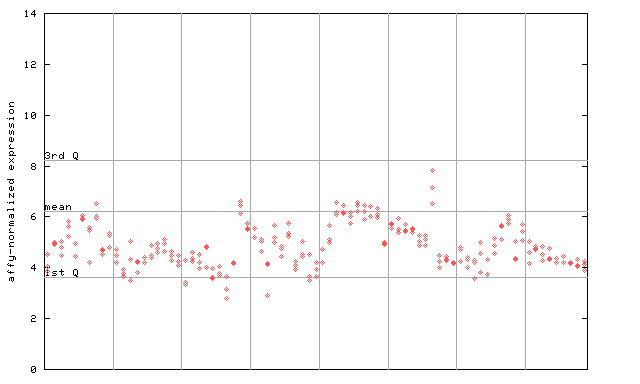
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
249907_at
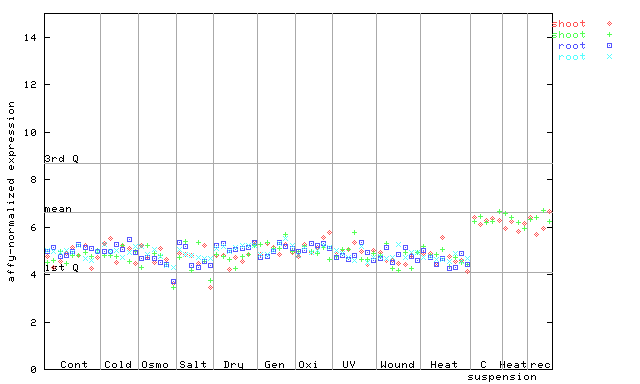
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
249907_at
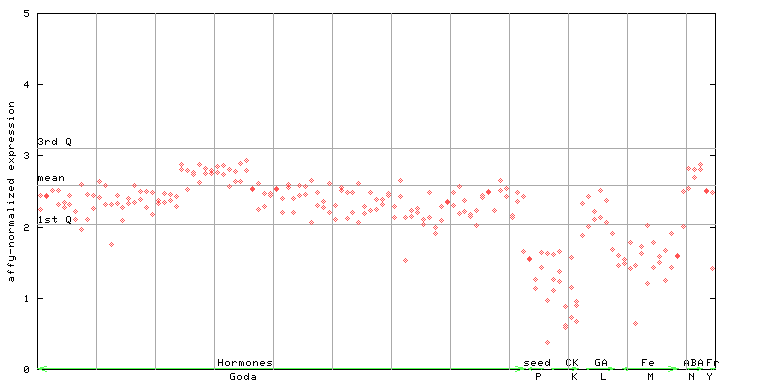
X axis is samples (xls file), and Y axis is log-expression.
|












