| functional annotation |
| Function |
temperature-induced lipocalin |


|
| GO BP |
|
GO:0030644 [list] [network] cellular chloride ion homeostasis
|
(1 genes)
|
IMP
|
 |
|
GO:1901562 [list] [network] response to paraquat
|
(4 genes)
|
IMP
|
 |
|
GO:0006883 [list] [network] cellular sodium ion homeostasis
|
(5 genes)
|
IMP
|
 |
|
GO:1902884 [list] [network] positive regulation of response to oxidative stress
|
(5 genes)
|
IMP
|
 |
|
GO:1901002 [list] [network] positive regulation of response to salt stress
|
(8 genes)
|
IMP
|
 |
|
GO:0050826 [list] [network] response to freezing
|
(28 genes)
|
IMP
|
 |
|
GO:0010286 [list] [network] heat acclimation
|
(53 genes)
|
IMP
|
 |
|
GO:0010431 [list] [network] seed maturation
|
(56 genes)
|
IMP
|
 |
|
GO:0042538 [list] [network] hyperosmotic salinity response
|
(57 genes)
|
IMP
|
 |
|
GO:0009644 [list] [network] response to high light intensity
|
(65 genes)
|
IMP
|
 |
|
GO:0000302 [list] [network] response to reactive oxygen species
|
(152 genes)
|
IBA
|
 |
|
GO:0009408 [list] [network] response to heat
|
(229 genes)
|
IEP
IMP
|
 |
|
GO:0071456 [list] [network] cellular response to hypoxia
|
(238 genes)
|
HEP
|
 |
|
GO:0009414 [list] [network] response to water deprivation
|
(361 genes)
|
IEP
|
 |
|
GO:0009409 [list] [network] response to cold
|
(411 genes)
|
IEP
|
 |
|
GO:0009416 [list] [network] response to light stimulus
|
(700 genes)
|
IMP
|
 |
|
GO:0006629 [list] [network] lipid metabolic process
|
(947 genes)
|
IBA
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0045735 [list] [network] nutrient reservoir activity
|
(55 genes)
|
IEA
|
 |
|
| KEGG |
|
|
| Protein |
NP_200615.1
|
| BLAST |
NP_200615.1
|
| Orthologous |
[Ortholog page]
LOC4329967 (osa)
LOC4345681 (osa)
LOC7489460 (ppo)
LOC11427119 (mtr)
LOC11436323 (mtr)
TIL (vvi)
LOC100272963 (zma)
LOC100286188 (zma)
TIL' (gma)
LOC100306133 (gma)
LOC100500030 (gma)
TIL (gma)
TIL (sly)
TIL' (sly)
LOC103856714 (bra)
|
Subcellular
localization
wolf |
|
nucl 4,
cyto 1,
mito 1,
extr 1,
pero 1,
cyto_pero 1
|
(predict for NP_200615.1)
|
|
Subcellular
localization
TargetP |
|
other 7
|
(predict for NP_200615.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath04141 |
Protein processing in endoplasmic reticulum |
4 |
 |
Genes directly connected with TIL on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 6.5 |
FBA6 |
Aldolase superfamily protein |
[detail] |
818220 |
| 5.6 |
AtCDC48B |
ATPase, AAA-type, CDC48 protein |
[detail] |
824489 |
| 5.4 |
AT4G13010 |
Oxidoreductase, zinc-binding dehydrogenase family protein |
[detail] |
826914 |
| 5.2 |
GolS1 |
galactinol synthase 1 |
[detail] |
819331 |
|
Coexpressed
gene list |
[Coexpressed gene list for TIL]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
247851_at
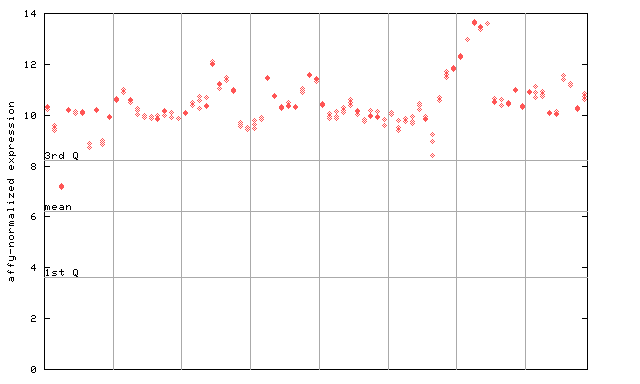
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
247851_at
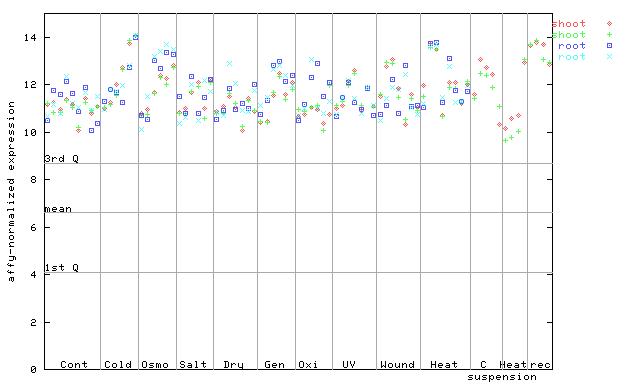
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
247851_at
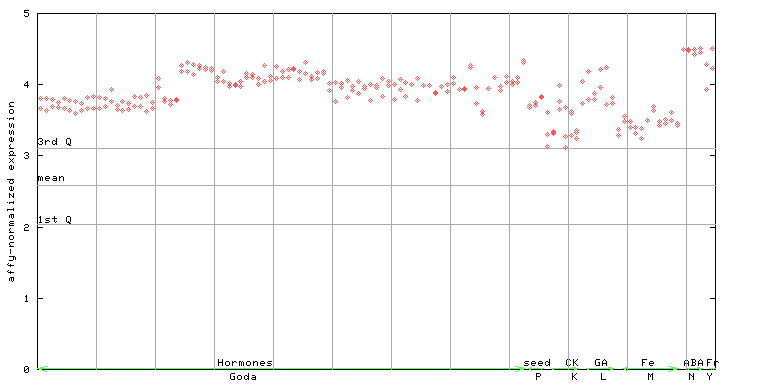
X axis is samples (xls file), and Y axis is log-expression.
|










