| functional annotation |
| Function |
cytochrome P450, family 78, subfamily A, polypeptide 5 |


|
| GO BP |
|
GO:0010338 [list] [network] leaf formation
|
(4 genes)
|
IGI
|
 |
|
GO:0040009 [list] [network] regulation of growth rate
|
(4 genes)
|
IMP
|
 |
|
GO:0046622 [list] [network] positive regulation of organ growth
|
(6 genes)
|
IMP
|
 |
|
GO:0035265 [list] [network] organ growth
|
(12 genes)
|
IMP
|
 |
|
GO:0008284 [list] [network] positive regulation of cell population proliferation
|
(26 genes)
|
IMP
|
 |
|
GO:0010075 [list] [network] regulation of meristem growth
|
(54 genes)
|
IGI
|
 |
|
GO:0048437 [list] [network] floral organ development
|
(242 genes)
|
IMP
|
 |
|
GO:0055114 [list] [network] oxidation-reduction process
|
(1468 genes)
|
IEA
|
 |
|
| GO CC |
|
GO:0005783 [list] [network] endoplasmic reticulum
|
(877 genes)
|
IDA
|
 |
|
GO:0016021 [list] [network] integral component of membrane
|
(4803 genes)
|
IEA
|
 |
|
GO:0009507 [list] [network] chloroplast
|
(5095 genes)
|
ISM
|
 |
|
| GO MF |
|
GO:0016709 [list] [network] oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen
|
(187 genes)
|
IBA
|
 |
|
GO:0005506 [list] [network] iron ion binding
|
(281 genes)
|
IEA
|
 |
|
GO:0020037 [list] [network] heme binding
|
(328 genes)
|
IEA
|
 |
|
| KEGG |
|
|
| Protein |
NP_172827.1
|
| BLAST |
NP_172827.1
|
| Orthologous |
[Ortholog page]
CYP78A10 (ath)
LOC4333118 (osa)
LOC4343840 (osa)
LOC11432063 (mtr)
LOC100249034 (vvi)
LOC100253660 (vvi)
LOC100381532 (zma)
LOC100783602 (gma)
LOC100790231 (gma)
CYP78A10 (gma)
LOC100804462 (gma)
LOC100811042 (gma)
LOC101258933 (sly)
LOC103636404 (zma)
LOC103639048 (zma)
LOC103832266 (bra)
LOC103843038 (bra)
LOC103848883 (bra)
LOC103852882 (bra)
LOC103852884 (bra)
|
Subcellular
localization
wolf |
|
chlo 9
|
(predict for NP_172827.1)
|
|
Subcellular
localization
TargetP |
|
scret 4
|
(predict for NP_172827.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
Genes directly connected with CYP78A5 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 8.6 |
STM |
KNOX/ELK homeobox transcription factor |
[detail] |
842534 |
| 8.5 |
GRF5 |
growth-regulating factor 5 |
[detail] |
820609 |
| 8.5 |
AT1G62500 |
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein |
[detail] |
842547 |
| 7.6 |
MYB17 |
myb domain protein 17 |
[detail] |
825297 |
| 5.5 |
AT4G30250 |
P-loop containing nucleoside triphosphate hydrolases superfamily protein |
[detail] |
829148 |
| 5.4 |
AT2G24700 |
Transcriptional factor B3 family protein |
[detail] |
817006 |
| 4.2 |
AT4G31620 |
Transcriptional factor B3 family protein |
[detail] |
829290 |
| 4.1 |
AT2G19920 |
RNA-dependent RNA polymerase family protein |
[detail] |
816511 |
| 3.6 |
AT2G19910 |
RNA-dependent RNA polymerase family protein |
[detail] |
816510 |
|
Coexpressed
gene list |
[Coexpressed gene list for CYP78A5]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
256099_at
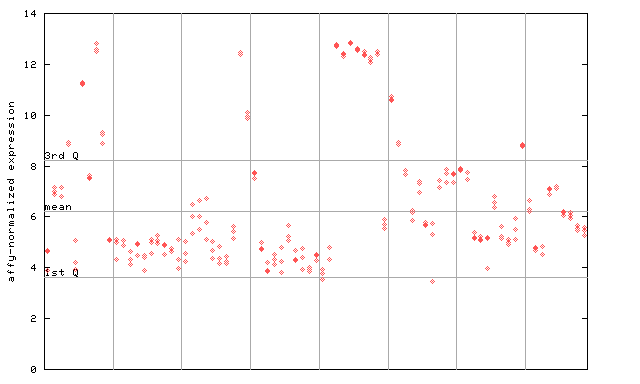
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
256099_at
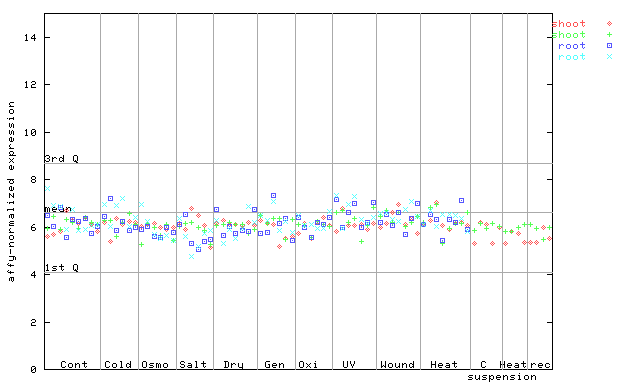
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
256099_at
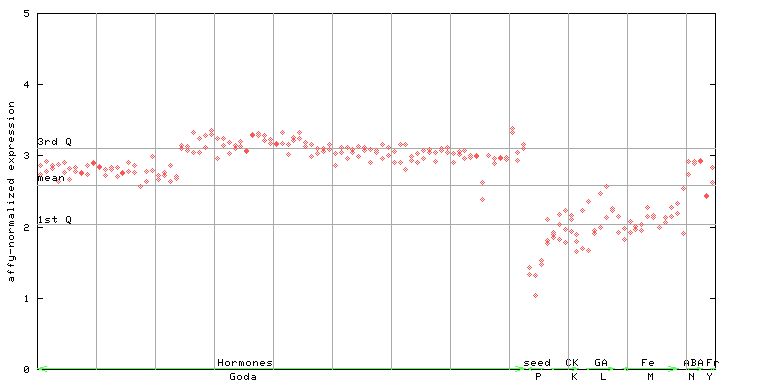
X axis is samples (xls file), and Y axis is log-expression.
|









