| functional annotation |
| Function |
WWE protein-protein interaction domain protein family |


|
| GO BP |
|
GO:0006809 [list] [network] nitric oxide biosynthetic process
|
(5 genes)
|
IMP
|
 |
|
GO:0000303 [list] [network] response to superoxide
|
(13 genes)
|
IMP
|
 |
|
GO:0010193 [list] [network] response to ozone
|
(36 genes)
|
IMP
|
 |
|
GO:0009816 [list] [network] defense response to bacterium, incompatible interaction
|
(43 genes)
|
IMP
|
 |
|
GO:0042542 [list] [network] response to hydrogen peroxide
|
(52 genes)
|
IMP
|
 |
|
GO:2000377 [list] [network] regulation of reactive oxygen species metabolic process
|
(54 genes)
|
IMP
|
 |
|
GO:0016032 [list] [network] viral process
|
(63 genes)
|
IEA
|
 |
|
GO:0009867 [list] [network] jasmonic acid mediated signaling pathway
|
(71 genes)
|
IMP
|
 |
|
GO:0010102 [list] [network] lateral root morphogenesis
|
(71 genes)
|
IMP
|
 |
|
GO:0012501 [list] [network] programmed cell death
|
(106 genes)
|
IMP
|
 |
|
GO:0009873 [list] [network] ethylene-activated signaling pathway
|
(191 genes)
|
IMP
|
 |
|
GO:0009723 [list] [network] response to ethylene
|
(297 genes)
|
IMP
|
 |
|
GO:0009414 [list] [network] response to water deprivation
|
(361 genes)
|
IMP
|
 |
|
GO:0006979 [list] [network] response to oxidative stress
|
(442 genes)
|
IGI
|
 |
|
GO:0009651 [list] [network] response to salt stress
|
(485 genes)
|
IMP
|
 |
|
GO:0009793 [list] [network] embryo development ending in seed dormancy
|
(547 genes)
|
IMP
|
 |
|
GO:0006970 [list] [network] response to osmotic stress
|
(563 genes)
|
IMP
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0003950 [list] [network] NAD+ ADP-ribosyltransferase activity
|
(9 genes)
|
IEA
|
 |
|
GO:0005515 [list] [network] protein binding
|
(4605 genes)
|
IPI
|
 |
|
| KEGG |
|
|
| Protein |
NP_001154388.1
NP_001320330.1
NP_001320331.1
NP_001320332.1
NP_564391.1
NP_849739.1
|
| BLAST |
NP_001154388.1
NP_001320330.1
NP_001320331.1
NP_001320332.1
NP_564391.1
NP_849739.1
|
| Orthologous |
[Ortholog page]
SRO1 (ath)
LOC4334829 (osa)
LOC4349501 (osa)
LOC7456095 (ppo)
LOC11405491 (mtr)
LOC11416324 (mtr)
LOC11424828 (mtr)
LOC25485596 (mtr)
LOC100245552 (vvi)
LOC100260925 (vvi)
LOC100797549 (gma)
LOC100800707 (gma)
LOC101027199 (zma)
LOC101254819 (sly)
LOC101256097 (sly)
LOC101265942 (sly)
LOC103625711 (zma)
LOC103637733 (zma)
LOC103833671 (bra)
LOC103840174 (bra)
LOC103853513 (bra)
LOC103865455 (bra)
LOC103867356 (bra)
LOC112421688 (mtr)
|
Subcellular
localization
wolf |
|
nucl 4,
cyto 4,
cyto_nucl 4
|
(predict for NP_001154388.1)
|
|
nucl 4,
cyto 4,
chlo 1,
pero 1,
cysk 1
|
(predict for NP_001320330.1)
|
|
nucl 4,
cyto 4,
chlo 1,
pero 1,
cysk 1
|
(predict for NP_001320331.1)
|
|
nucl 4,
cyto 4,
chlo 1,
pero 1,
cysk 1
|
(predict for NP_001320332.1)
|
|
nucl 4,
cyto 4,
chlo 1,
pero 1,
cysk 1
|
(predict for NP_564391.1)
|
|
nucl 4,
cyto 4,
chlo 1,
pero 1,
cysk 1
|
(predict for NP_849739.1)
|
|
Subcellular
localization
TargetP |
|
other 8
|
(predict for NP_001154388.1)
|
|
other 8
|
(predict for NP_001320330.1)
|
|
other 8
|
(predict for NP_001320331.1)
|
|
other 8
|
(predict for NP_001320332.1)
|
|
other 8
|
(predict for NP_564391.1)
|
|
other 8
|
(predict for NP_849739.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
Genes directly connected with RCD1 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 7.2 |
AT5G55600 |
Agenet and bromo-adjacent homology (BAH) domain-containing protein |
[detail] |
835654 |
| 6.7 |
SRO1 |
similar to RCD one 1 |
[detail] |
818116 |
| 6.3 |
AT5G16880 |
Target of Myb protein 1 |
[detail] |
831551 |
| 5.0 |
CIPK23 |
CBL-interacting protein kinase 23 |
[detail] |
839907 |
|
Coexpressed
gene list |
[Coexpressed gene list for RCD1]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
245796_at
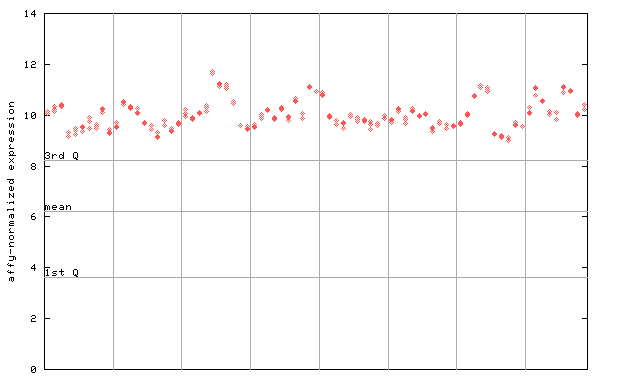
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
245796_at
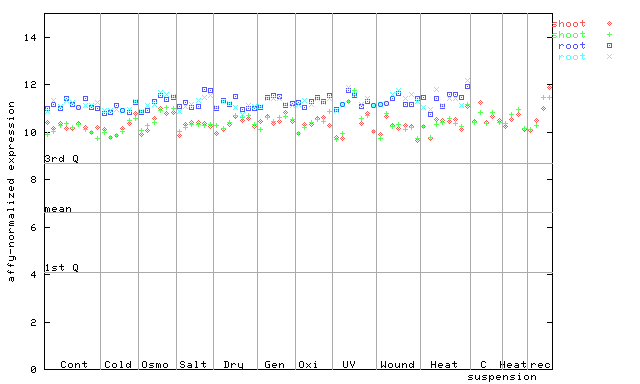
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
245796_at
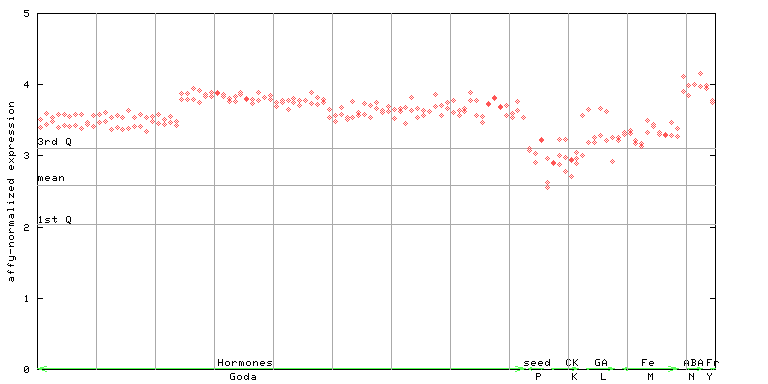
X axis is samples (xls file), and Y axis is log-expression.
|














