[←][→] ath
| functional annotation | ||||||||||||||||||||||||||||||||||||||
| Function | alpha dioxygenase |




|
||||||||||||||||||||||||||||||||||||
| GO BP |
|
|||||||||||||||||||||||||||||||||||||
| GO CC |
|
|||||||||||||||||||||||||||||||||||||
| GO MF |
|
|||||||||||||||||||||||||||||||||||||
| KEGG | ath00592 [list] [network] alpha-Linolenic acid metabolism (43 genes) |  |
||||||||||||||||||||||||||||||||||||
| Protein | NP_001185393.1 NP_001322742.1 NP_001322743.1 NP_177509.1 | |||||||||||||||||||||||||||||||||||||
| BLAST | NP_001185393.1 NP_001322742.1 NP_001322743.1 NP_177509.1 | |||||||||||||||||||||||||||||||||||||
| Orthologous | [Ortholog page] alpha-DOX2 (sly) LOC7486739 (ppo) LOC11412210 (mtr) LOC103830737 (bra) LOC103831843 (bra) LOC103852857 (bra) LOC112323590 (ppo) | |||||||||||||||||||||||||||||||||||||
| Subcellular localization wolf |
|
|||||||||||||||||||||||||||||||||||||
| Subcellular localization TargetP |
|
|||||||||||||||||||||||||||||||||||||
| Gene coexpression | ||||||||||||||||||||||||||||||||||||||
| Network*for coexpressed genes |
|
|||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Coexpressed gene list |
[Coexpressed gene list for ALPHA DOX2] | |||||||||||||||||||||||||||||||||||||
| Gene expression | ||||||||||||||||||||||||||||||||||||||
| All samples | [Expression pattern for all samples] | |||||||||||||||||||||||||||||||||||||
| AtGenExpress* (Development) |
260060_at
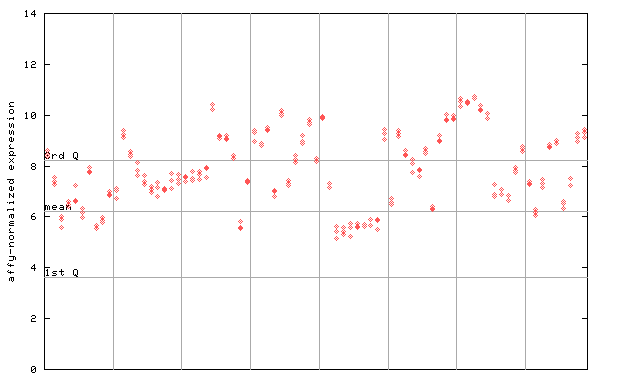
X axis is samples (pdf file), and Y axis is log2-expression. |
|||||||||||||||||||||||||||||||||||||
| AtGenExpress* (Stress) |
260060_at
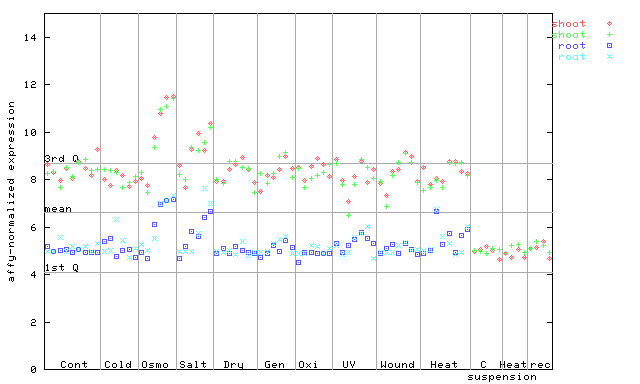
X axis is samples (pdf file), and Y axis is log2-expression. |
|||||||||||||||||||||||||||||||||||||
| AtGenExpress* (Hormone) |
260060_at
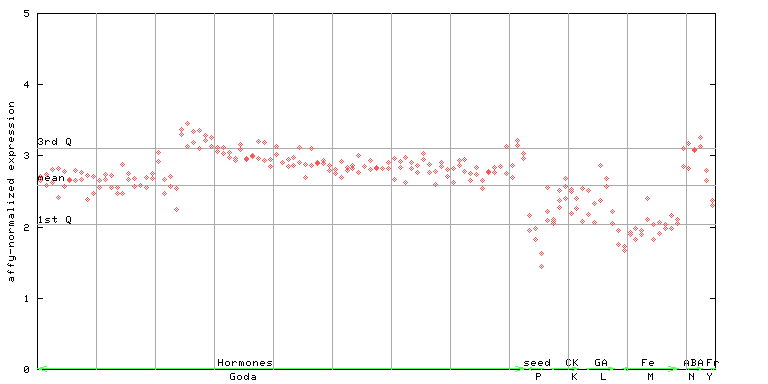
X axis is samples (xls file), and Y axis is log-expression. |
|||||||||||||||||||||||||||||||||||||
| Link to other DBs | ||
| Entrez Gene ID | 843703 |


|
| Refseq ID (protein) | NP_001185393.1 |  |
| NP_001322742.1 |  |
|
| NP_001322743.1 |  |
|
| NP_177509.1 |  |
|
The preparation time of this page was 0.1 [sec].



