| functional annotation |
| Function |
NAD(P)-linked oxidoreductase superfamily protein |


|
| GO BP |
|
GO:0009636 [list] [network] response to toxic substance
|
(192 genes)
|
IEA
|
 |
|
GO:0046686 [list] [network] response to cadmium ion
|
(346 genes)
|
IEP
|
 |
|
GO:0009414 [list] [network] response to water deprivation
|
(361 genes)
|
IEP
|
 |
|
GO:0009409 [list] [network] response to cold
|
(411 genes)
|
IEP
|
 |
|
GO:0009651 [list] [network] response to salt stress
|
(485 genes)
|
IEP
|
 |
|
GO:0055114 [list] [network] oxidation-reduction process
|
(1468 genes)
|
IDA
|
 |
|
| GO CC |
|
| GO MF |
|
| KEGG |
|
|
| Protein |
NP_001078019.1
NP_001118465.1
NP_001324984.1
NP_001324985.1
NP_565871.1
NP_973626.2
NP_973627.1
|
| BLAST |
NP_001078019.1
NP_001118465.1
NP_001324984.1
NP_001324985.1
NP_565871.1
NP_973626.2
NP_973627.1
|
| Orthologous |
[Ortholog page]
LOC101251566 (sly)
LOC103867164 (bra)
|
Subcellular
localization
wolf |
|
cyto 3,
chlo 3,
nucl 3,
cyto_E.R. 2
|
(predict for NP_001078019.1)
|
|
cyto 3,
chlo 3,
nucl 3,
cyto_E.R. 2
|
(predict for NP_001118465.1)
|
|
nucl 4,
cyto 3,
cyto_E.R. 2,
mito 1,
chlo 1,
extr 1,
cyto_mito 1,
mito_plas 1
|
(predict for NP_001324984.1)
|
|
nucl 4,
cyto 3,
cyto_E.R. 2,
mito 1,
chlo 1,
extr 1,
cyto_mito 1,
mito_plas 1
|
(predict for NP_001324985.1)
|
|
cyto 5,
cyto_E.R. 3,
nucl 2,
chlo 1,
cysk_nucl 1
|
(predict for NP_565871.1)
|
|
cyto 5,
cyto_E.R. 3,
nucl 2,
chlo 1,
cysk_nucl 1
|
(predict for NP_973626.2)
|
|
cyto 3,
chlo 3,
nucl 3,
cyto_E.R. 2
|
(predict for NP_973627.1)
|
|
Subcellular
localization
TargetP |
|
other 5,
scret 3
|
(predict for NP_001078019.1)
|
|
other 5,
scret 3
|
(predict for NP_001118465.1)
|
|
other 6,
scret 3
|
(predict for NP_001324984.1)
|
|
other 6,
scret 3
|
(predict for NP_001324985.1)
|
|
other 5,
scret 3
|
(predict for NP_565871.1)
|
|
other 5,
scret 3
|
(predict for NP_973626.2)
|
|
other 5,
scret 3
|
(predict for NP_973627.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath00940 |
Phenylpropanoid biosynthesis |
3 |
 |
| ath01230 |
Biosynthesis of amino acids |
3 |
 |
| ath00460 |
Cyanoamino acid metabolism |
2 |
 |
| ath00500 |
Starch and sucrose metabolism |
2 |
 |
| ath00350 |
Tyrosine metabolism |
2 |
 |
Genes directly connected with AKR4C8 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 13.0 |
BGLU11 |
beta glucosidase 11 |
[detail] |
839435 |
| 12.5 |
ChlAKR |
NAD(P)-linked oxidoreductase superfamily protein |
[detail] |
818354 |
| 7.5 |
GSTZ1 |
glutathione S-transferase zeta 1 |
[detail] |
814770 |
| 6.2 |
BGLU10 |
beta glucosidase 10 |
[detail] |
828896 |
| 6.0 |
AT5G36160 |
Tyrosine transaminase family protein |
[detail] |
833613 |
| 5.1 |
AT3G04000 |
NAD(P)-binding Rossmann-fold superfamily protein |
[detail] |
819555 |
| 4.6 |
THA2 |
threonine aldolase 2 |
[detail] |
819608 |
|
Coexpressed
gene list |
[Coexpressed gene list for AKR4C8]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
267181_at
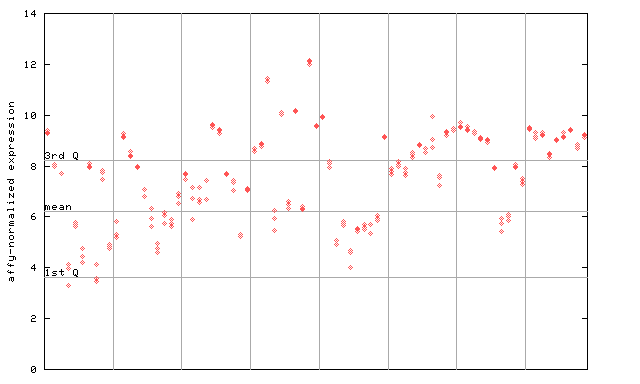
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
267181_at
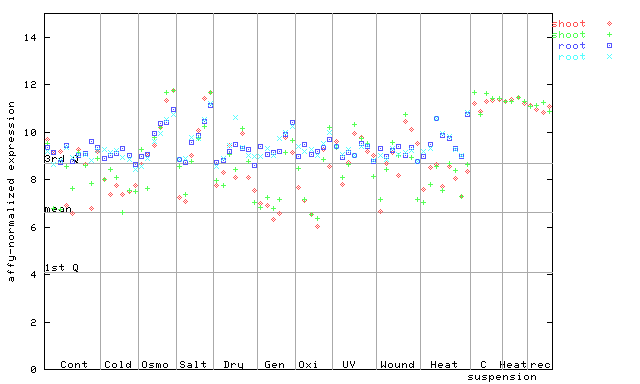
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
267181_at
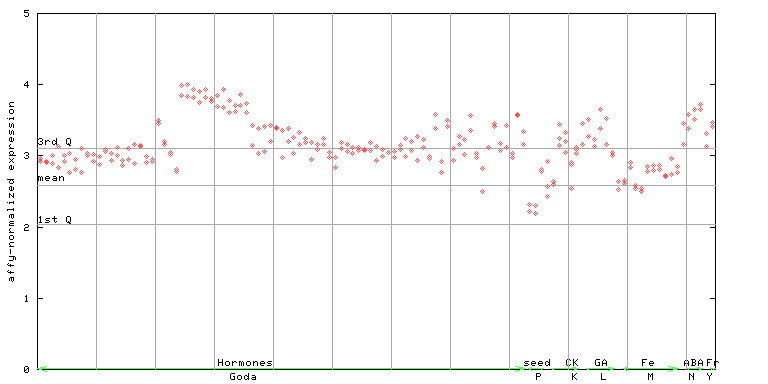
X axis is samples (xls file), and Y axis is log-expression.
|




















