| functional annotation |
| Function |
Glycosyl hydrolase superfamily protein |


|
| GO BP |
|
GO:0042344 [list] [network] indole glucosinolate catabolic process
|
(5 genes)
|
IMP
|
 |
|
GO:0052544 [list] [network] defense response by callose deposition in cell wall
|
(16 genes)
|
IMP
|
 |
|
GO:0009682 [list] [network] induced systemic resistance
|
(32 genes)
|
IMP
|
 |
|
GO:0019762 [list] [network] glucosinolate catabolic process
|
(34 genes)
|
IBA
|
 |
|
GO:0009817 [list] [network] defense response to fungus, incompatible interaction
|
(50 genes)
|
IMP
|
 |
|
GO:0019760 [list] [network] glucosinolate metabolic process
|
(111 genes)
|
IMP
|
 |
|
GO:0042742 [list] [network] defense response to bacterium
|
(399 genes)
|
IMP
|
 |
|
GO:0009617 [list] [network] response to bacterium
|
(479 genes)
|
IMP
|
 |
|
GO:0009651 [list] [network] response to salt stress
|
(485 genes)
|
IBA
|
 |
|
GO:0005975 [list] [network] carbohydrate metabolic process
|
(995 genes)
|
IEA
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0019137 [list] [network] thioglucosidase activity
|
(8 genes)
|
IDA
|
 |
|
GO:0102483 [list] [network] scopolin beta-glucosidase activity
|
(42 genes)
|
IEA
|
 |
|
GO:0008422 [list] [network] beta-glucosidase activity
|
(77 genes)
|
IBA
|
 |
|
| KEGG |
ath00460 [list] [network] Cyanoamino acid metabolism (69 genes) |
 |
| ath00500 [list] [network] Starch and sucrose metabolism (165 genes) |
 |
| ath00940 [list] [network] Phenylpropanoid biosynthesis (166 genes) |
 |
| Protein |
NP_181977.1
|
| BLAST |
NP_181977.1
|
| Orthologous |
[Ortholog page]
LOC103866804 (bra)
|
Subcellular
localization
wolf |
|
chlo 3,
nucl 3,
plas 1,
golg 1,
chlo_mito 1,
cysk_nucl 1,
golg_plas 1
|
(predict for NP_181977.1)
|
|
Subcellular
localization
TargetP |
|
mito 6,
other 3
|
(predict for NP_181977.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath00940 |
Phenylpropanoid biosynthesis |
2 |
 |
Genes directly connected with PEN2 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 10.9 |
AT1G55450 |
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein |
[detail] |
841992 |
| 9.7 |
CAD1 |
phytochelatin synthase 1 (PCS1) |
[detail] |
834430 |
| 9.4 |
PEN3 |
ABC-2 and Plant PDR ABC-type transporter family protein |
[detail] |
842281 |
| 9.3 |
pEARLI4 |
phospholipase-like protein (PEARLI 4) family protein |
[detail] |
816630 |
| 8.4 |
AT3G19010 |
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein |
[detail] |
821434 |
| 7.6 |
MLO2 |
Seven transmembrane MLO family protein |
[detail] |
837673 |
| 7.3 |
AT1G53430 |
Leucine-rich repeat transmembrane protein kinase |
[detail] |
841778 |
| 7.1 |
AT1G30755 |
elongation factor G, putative (DUF668) |
[detail] |
839956 |
| 6.9 |
CRK19 |
cysteine-rich RLK (RECEPTOR-like protein kinase) 19 |
[detail] |
828426 |
| 6.9 |
AT5G20050 |
Protein kinase superfamily protein |
[detail] |
832127 |
| 6.9 |
CYP71B7 |
cytochrome P450, family 71 subfamily B, polypeptide 7 |
[detail] |
837868 |
| 6.6 |
AT1G18390 |
Serine/Threonine kinase family catalytic domain protein |
[detail] |
838420 |
| 6.3 |
AT5G49760 |
Leucine-rich repeat protein kinase family protein |
[detail] |
835039 |
| 6.0 |
WAKL6 |
wall associated kinase-like 6 |
[detail] |
838180 |
| 5.8 |
AT1G51805 |
Leucine-rich repeat protein kinase family protein |
[detail] |
841607 |
| 5.4 |
SHB1 |
EXS (ERD1/XPR1/SYG1) family protein |
[detail] |
828638 |
| 5.1 |
UTR2 |
UDP-galactose transporter 2 |
[detail] |
828400 |
| 4.9 |
LYK4 |
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein |
[detail] |
816909 |
| 4.8 |
MPK5 |
MAP kinase 5 |
[detail] |
826735 |
| 4.7 |
ALDH2C4 |
aldehyde dehydrogenase 2C4 |
[detail] |
822042 |
|
Coexpressed
gene list |
[Coexpressed gene list for PEN2]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
267392_at
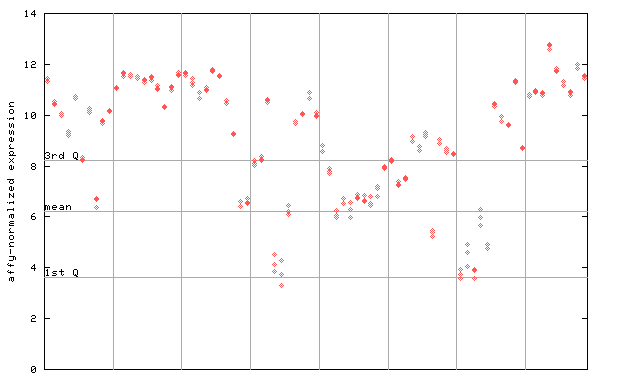
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
267392_at
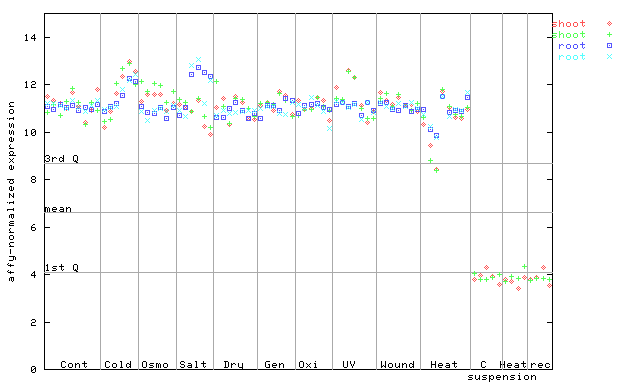
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
267392_at
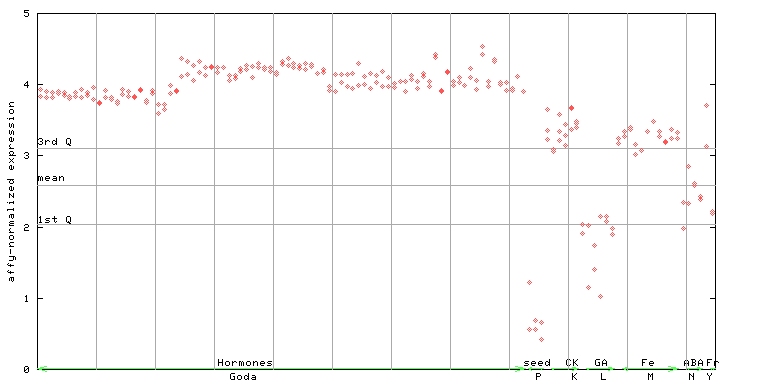
X axis is samples (xls file), and Y axis is log-expression.
|













