[←][→] ath
| functional annotation | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Function | cystathionine beta-lyase |

 Plant GARDEN Plant GARDEN JBrowse
Plant GARDEN Plant GARDEN JBrowse
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO BP |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO CC |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO MF |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG | ath00270 [list] [network] Cysteine and methionine metabolism (124 genes) |  |
|||||||||||||||||||||||||||||||||||||||||||||||||||||
| ath00450 [list] [network] Selenocompound metabolism (18 genes) |  |
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ath01230 [list] [network] Biosynthesis of amino acids (244 genes) |  |
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein | NP_001326867.1 NP_001326868.1 NP_191264.1 NP_850712.1 NP_850713.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BLAST | NP_001326867.1 NP_001326868.1 NP_191264.1 NP_850712.1 NP_850713.1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Orthologous | [Ortholog page] LOC4340283 (osa) LOC18106110 (ppo) LOC25483994 (mtr) LOC100786470 (gma) LOC100812354 (gma) LOC101265082 (sly) LOC103841611 (bra) LOC103863064 (bra) LOC107275791 (osa) LOC123150324 (tae) LOC123160296 (tae) LOC123166424 (tae) LOC123409968 (hvu) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Subcellular localization wolf |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Subcellular localization TargetP |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene coexpression | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Network*for coexpressed genes |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Coexpressed gene list |
[Coexpressed gene list for CBL] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene expression | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
| All samples | [Expression pattern for all samples] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| AtGenExpress* (Development) |
251666_at
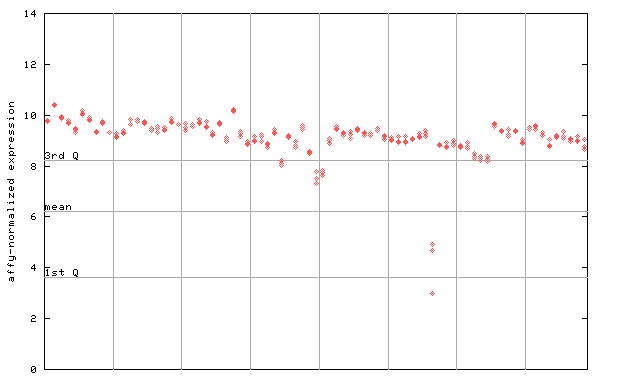
X axis is samples (pdf file), and Y axis is log2-expression. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| AtGenExpress* (Stress) |
251666_at
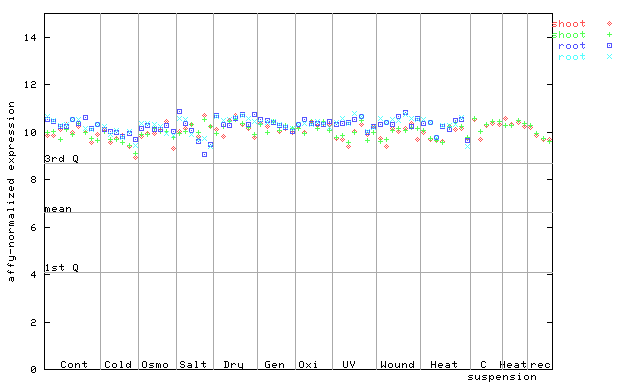
X axis is samples (pdf file), and Y axis is log2-expression. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| AtGenExpress* (Hormone) |
251666_at
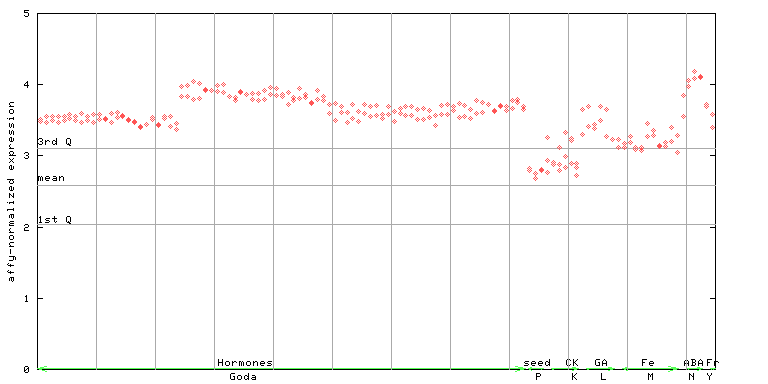
X axis is samples (xls file), and Y axis is log-expression. |
||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Link to other DBs | ||
| Entrez Gene ID | 824872 |


|
| Refseq ID (protein) | NP_001326867.1 |  |
| NP_001326868.1 |  |
|
| NP_191264.1 |  |
|
| NP_850712.1 |  |
|
| NP_850713.1 |  |
|
The preparation time of this page was 0.1 [sec].






