| functional annotation |
| Function |
pyruvate carrier-like protein |


|
| GO BP |
|
GO:1901017 [list] [network] negative regulation of potassium ion transmembrane transporter activity
|
(1 genes)
|
IMP
|
 |
|
GO:0010360 [list] [network] negative regulation of anion channel activity
|
(3 genes)
|
IMP
|
 |
|
GO:0006850 [list] [network] mitochondrial pyruvate transmembrane transport
|
(4 genes)
|
IBA
IMP
|
 |
|
GO:2000070 [list] [network] regulation of response to water deprivation
|
(31 genes)
|
IMP
|
 |
|
| GO CC |
|
GO:0031305 [list] [network] integral component of mitochondrial inner membrane
|
(23 genes)
|
IBA
|
 |
|
GO:0005739 [list] [network] mitochondrion
|
(4405 genes)
|
IDA
|
 |
|
GO:0009507 [list] [network] chloroplast
|
(5095 genes)
|
ISM
|
 |
|
| GO MF |
|
GO:0050833 [list] [network] pyruvate transmembrane transporter activity
|
(4 genes)
|
IBA
|
 |
|
GO:0005515 [list] [network] protein binding
|
(4605 genes)
|
IPI
|
 |
|
| KEGG |
|
|
| Protein |
NP_001031587.1
NP_001329913.1
NP_567306.1
|
| BLAST |
NP_001031587.1
NP_001329913.1
NP_567306.1
|
| Orthologous |
|
Subcellular
localization
wolf |
|
cyto 6,
chlo 3
|
(predict for NP_001031587.1)
|
|
extr 4,
cyto 2,
chlo 1,
nucl 1,
mito 1,
cyto_pero 1,
mito_plas 1,
cyto_E.R. 1
|
(predict for NP_001329913.1)
|
|
cyto 3,
chlo 2,
extr 2,
mito 1,
cyto_nucl 1
|
(predict for NP_567306.1)
|
|
Subcellular
localization
TargetP |
|
other 6,
mito 4
|
(predict for NP_001031587.1)
|
|
other 5,
mito 3
|
(predict for NP_001329913.1)
|
|
other 6,
mito 4
|
(predict for NP_567306.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath01200 |
Carbon metabolism |
5 |
 |
| ath00020 |
Citrate cycle (TCA cycle) |
2 |
 |
| ath01210 |
2-Oxocarboxylic acid metabolism |
2 |
 |
| ath01230 |
Biosynthesis of amino acids |
2 |
 |
| ath00260 |
Glycine, serine and threonine metabolism |
2 |
 |
Genes directly connected with AT4G05590 on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 4.8 |
AT4G10300 |
RmlC-like cupins superfamily protein |
[detail] |
826622 |
| 4.7 |
ABCI17 |
non-intrinsic ABC protein 3 |
[detail] |
843122 |
| 4.6 |
AT3G28950 |
AIG2-like (avirulence induced gene) family protein |
[detail] |
822532 |
|
Coexpressed
gene list |
[Coexpressed gene list for AT4G05590]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
255243_at
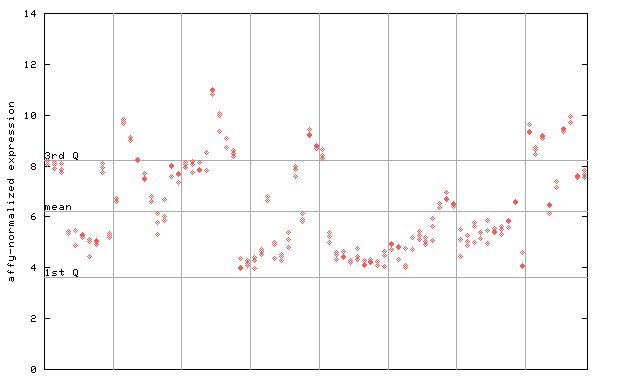
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
255243_at
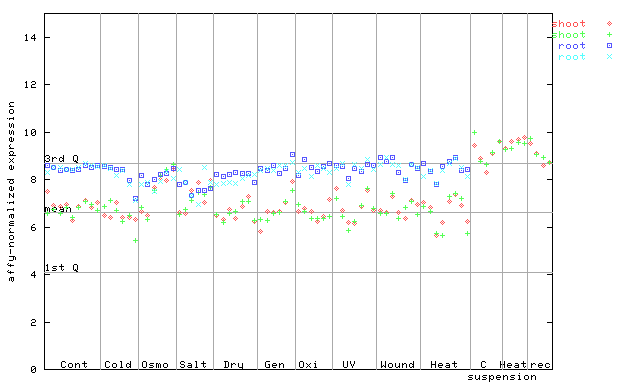
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
255243_at
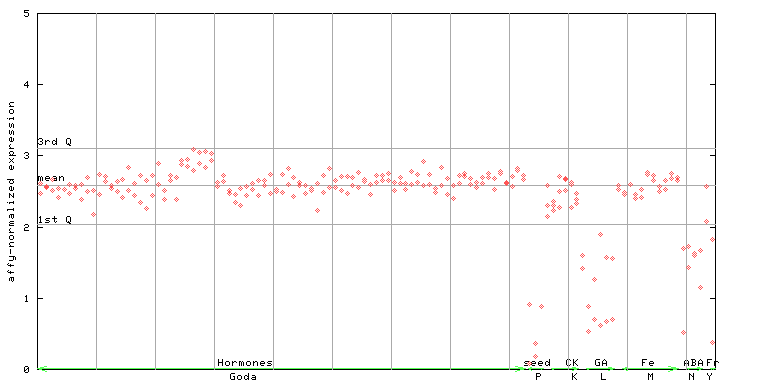
X axis is samples (xls file), and Y axis is log-expression.
|
















