| functional annotation |
| Function |
uncharacterized protein |




|
| GO BP |
|
GO:0009684 [list] [network] indoleacetic acid biosynthetic process
|
(11 genes)
|
IMP
|
 |
|
GO:0009641 [list] [network] shade avoidance
|
(17 genes)
|
IMP
|
 |
|
GO:0010588 [list] [network] cotyledon vascular tissue pattern formation
|
(19 genes)
|
IGI
|
 |
|
GO:0080022 [list] [network] primary root development
|
(36 genes)
|
IGI
|
 |
|
GO:0009958 [list] [network] positive gravitropism
|
(39 genes)
|
IMP
|
 |
|
GO:0010078 [list] [network] maintenance of root meristem identity
|
(40 genes)
|
IGI
|
 |
|
GO:0048825 [list] [network] cotyledon development
|
(59 genes)
|
IGI
|
 |
|
GO:0009734 [list] [network] auxin-activated signaling pathway
|
(65 genes)
|
NAS
|
 |
|
GO:0009926 [list] [network] auxin polar transport
|
(69 genes)
|
IMP
|
 |
|
GO:0048467 [list] [network] gynoecium development
|
(101 genes)
|
IGI
|
 |
|
GO:0010087 [list] [network] phloem or xylem histogenesis
|
(158 genes)
|
IGI
|
 |
|
GO:0009723 [list] [network] response to ethylene
|
(241 genes)
|
IEP
IMP
|
 |
|
GO:0048366 [list] [network] leaf development
|
(465 genes)
|
IMP
|
 |
|
GO:0009908 [list] [network] flower development
|
(492 genes)
|
IGI
|
 |
|
GO:0009793 [list] [network] embryo development ending in seed dormancy
|
(697 genes)
|
IGI
|
 |
|
GO:0042742 [list] [network] defense response to bacterium
|
(980 genes)
|
IGI
|
 |
|
GO:0048364 [list] [network] root development
|
(1192 genes)
|
IMP
|
 |
|
GO:0048367 [list] [network] shoot system development
|
(1318 genes)
|
IGI
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0047312 [list] [network] L-phenylalanine:pyruvate aminotransferase activity
|
(1 genes)
|
IDA
|
 |
|
GO:0050048 [list] [network] L-leucine:2-oxoglutarate aminotransferase activity
|
(1 genes)
|
IDA
|
 |
|
GO:0080098 [list] [network] L-tyrosine:pyruvate aminotransferase activity
|
(1 genes)
|
IDA
|
 |
|
GO:0080099 [list] [network] L-methionine:2-oxoglutarate aminotransferase activity
|
(1 genes)
|
IDA
|
 |
|
GO:0080100 [list] [network] L-glutamine:2-oxoglutarate aminotransferase activity
|
(1 genes)
|
IDA
|
 |
|
GO:0080130 [list] [network] L-phenylalanine:2-oxoglutarate aminotransferase activity
|
(1 genes)
|
IDA
|
 |
|
GO:0080097 [list] [network] L-tryptophan:pyruvate aminotransferase activity
|
(3 genes)
|
IDA
|
 |
|
GO:0050362 [list] [network] L-tryptophan:2-oxoglutarate aminotransferase activity
|
(4 genes)
|
IDA
|
 |
|
GO:0004021 [list] [network] L-alanine:2-oxoglutarate aminotransferase activity
|
(5 genes)
|
IDA
|
 |
|
GO:0004838 [list] [network] L-tyrosine:2-oxoglutarate aminotransferase activity
|
(5 genes)
|
IDA
|
 |
|
GO:0030170 [list] [network] pyridoxal phosphate binding
|
(26 genes)
|
IDA
|
 |
|
GO:0005515 [list] [network] protein binding
|
(5066 genes)
|
IPI
|
 |
|
| KEGG |
ath00380 [list] [network] Tryptophan metabolism (64 genes) |
 |
| Protein |
NP_177213.1
|
| BLAST |
NP_177213.1
|
| Orthologous |
[Ortholog page]
TAR2 (ath)
TAR1 (ath)
LOC4325198 (osa)
LOC4337930 (osa)
LOC7456901 (ppo)
LOC7492386 (ppo)
LOC11410630 (mtr)
LOC11415573 (mtr)
LOC11439449 (mtr)
LOC25493735 (mtr)
LOC100777098 (gma)
LOC100801447 (gma)
LOC100803246 (gma)
LOC100807013 (gma)
LOC100818522 (gma)
LOC100820582 (gma)
TAA1 (sly)
LOC101253797 (sly)
LOC101262607 (sly)
LOC103831591 (bra)
LOC103840774 (bra)
LOC103840776 (bra)
LOC103852664 (bra)
LOC103862382 (bra)
LOC112324299 (ppo)
LOC112328448 (ppo)
LOC123053434 (tae)
LOC123068780 (tae)
LOC123077285 (tae)
LOC123083178 (tae)
LOC123083188 (tae)
LOC123083217 (tae)
LOC123083248 (tae)
LOC123105817 (tae)
LOC123114279 (tae)
LOC123114319 (tae)
LOC123130433 (tae)
LOC123151276 (tae)
LOC123169461 (tae)
LOC123176510 (tae)
LOC123180651 (tae)
LOC123407837 (hvu)
LOC123414958 (hvu)
LOC123442807 (hvu)
|
Subcellular
localization
wolf |
|
cyto 7,
chlo 1,
nucl 1,
plas 1,
cysk 1,
cysk_nucl 1,
nucl_plas 1,
cysk_plas 1
|
(predict for NP_177213.1)
|
|
Subcellular
localization
TargetP |
|
other 8
|
(predict for NP_177213.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
Genes directly connected with TAA1 on the network
| coex z* |
Locus |
Function* |
CoexViewer |
Entrez Gene ID* |
| 5.5 |
WAG1 |
WAG 1 |
[detail] |
841807 |
| 5.4 |
ACR4 |
ACT domain repeat 4 |
[detail] |
843236 |
| 4.6 |
AT1G75710 |
C2H2-like zinc finger protein |
[detail] |
843905 |
|
Coexpressed
gene list |
[Coexpressed gene list for TAA1]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
260364_at
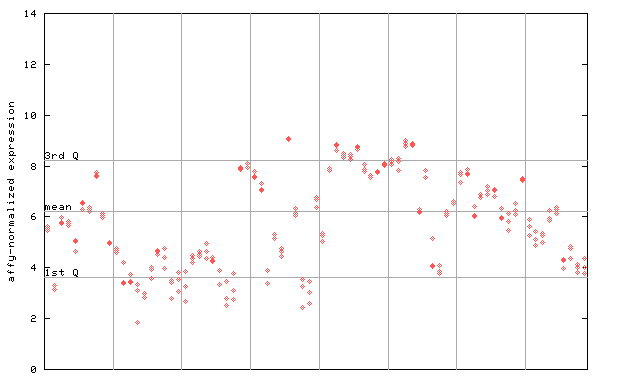
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
260364_at
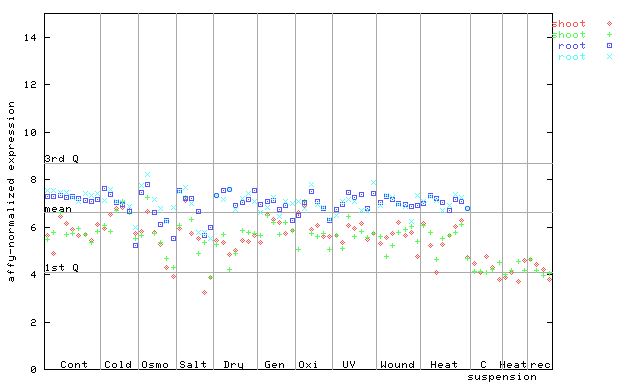
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
260364_at
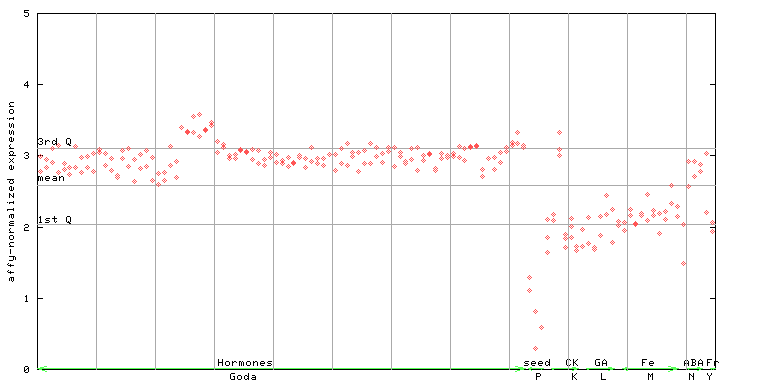
X axis is samples (xls file), and Y axis is log-expression.
|












