| functional annotation |
| Function |
phytochrome B |




|
| GO BP |
|
GO:0017012 [list] [network] protein-phytochromobilin linkage
|
(1 genes)
|
IDA
|
 |
|
GO:0010202 [list] [network] response to low fluence red light stimulus
|
(2 genes)
|
IMP
|
 |
|
GO:0010617 [list] [network] circadian regulation of calcium ion oscillation
|
(3 genes)
|
IMP
|
 |
|
GO:0009584 [list] [network] detection of visible light
|
(5 genes)
|
IEA
|
 |
|
GO:0010148 [list] [network] transpiration
|
(6 genes)
|
IMP
|
 |
|
GO:0010244 [list] [network] response to low fluence blue light stimulus by blue low-fluence system
|
(9 genes)
|
IDA
|
 |
|
GO:0010161 [list] [network] red light signaling pathway
|
(10 genes)
|
IMP
|
 |
|
GO:0009649 [list] [network] entrainment of circadian clock
|
(12 genes)
|
IMP
|
 |
|
GO:0009687 [list] [network] abscisic acid metabolic process
|
(21 genes)
|
IMP
|
 |
|
GO:0009638 [list] [network] phototropism
|
(30 genes)
|
IMP
|
 |
|
GO:0010218 [list] [network] response to far red light
|
(48 genes)
|
IMP
|
 |
|
GO:0010374 [list] [network] stomatal complex development
|
(75 genes)
|
IMP
|
 |
|
GO:0009640 [list] [network] photomorphogenesis
|
(80 genes)
|
IMP
|
 |
|
GO:0010029 [list] [network] regulation of seed germination
|
(93 genes)
|
IMP
|
 |
|
GO:2000028 [list] [network] regulation of photoperiodism, flowering
|
(94 genes)
|
IMP
|
 |
|
GO:0009630 [list] [network] gravitropism
|
(103 genes)
|
IMP
|
 |
|
GO:0009867 [list] [network] jasmonic acid mediated signaling pathway
|
(147 genes)
|
IMP
|
 |
|
GO:0015979 [list] [network] photosynthesis
|
(167 genes)
|
IMP
|
 |
|
GO:0045892 [list] [network] negative regulation of DNA-templated transcription
|
(205 genes)
|
IDA
|
 |
|
GO:0006325 [list] [network] chromatin organization
|
(210 genes)
|
IMP
|
 |
|
GO:0009409 [list] [network] response to cold
|
(472 genes)
|
IMP
|
 |
|
GO:0031347 [list] [network] regulation of defense response
|
(749 genes)
|
IMP
|
 |
|
GO:0009266 [list] [network] response to temperature stimulus
|
(843 genes)
|
IMP
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0031517 [list] [network] red light photoreceptor activity
|
(1 genes)
|
IDA
|
 |
|
GO:0031516 [list] [network] far-red light photoreceptor activity
|
(2 genes)
|
IDA
|
 |
|
GO:0009883 [list] [network] red or far-red light photoreceptor activity
|
(3 genes)
|
IMP
|
 |
|
GO:0000155 [list] [network] phosphorelay sensor kinase activity
|
(14 genes)
|
IEA
|
 |
|
GO:1990841 [list] [network] promoter-specific chromatin binding
|
(15 genes)
|
IDA
|
 |
|
GO:0004673 [list] [network] protein histidine kinase activity
|
(17 genes)
|
ISS
|
 |
|
GO:0042803 [list] [network] protein homodimerization activity
|
(227 genes)
|
IDA
|
 |
|
GO:0042802 [list] [network] identical protein binding
|
(404 genes)
|
IPI
|
 |
|
GO:0043565 [list] [network] sequence-specific DNA binding
|
(890 genes)
|
IDA
|
 |
|
GO:0005515 [list] [network] protein binding
|
(5066 genes)
|
IPI
|
 |
|
| KEGG |
ath04712 [list] [network] Circadian rhythm - plant (39 genes) |
 |
| Protein |
NP_001325249.1
NP_179469.1
|
| BLAST |
NP_001325249.1
NP_179469.1
|
| Orthologous |
[Ortholog page]
PHYD (ath)
PHYE (ath)
LOC4332623 (osa)
LOC7485825 (ppo)
LOC7485902 (ppo)
LOC11417317 (mtr)
LOC11420025 (mtr)
LOC100137004 (tae)
LOC100794865 (gma)
PHYB (gma)
PHYE2 (gma)
PHYE1 (gma)
PHYE (sly)
PHYB1 (sly)
PHYB2 (sly)
LOC103856075 (bra)
LOC103869720 (bra)
LOC123092340 (tae)
LOC123097663 (tae)
LOC123448812 (hvu)
|
Subcellular
localization
wolf |
|
nucl 3,
plas 2,
vacu 1,
cysk_plas 1,
E.R._plas 1,
cyto_plas 1
|
(predict for NP_001325249.1)
|
|
nucl 5,
plas 2,
chlo 1,
cyto_plas 1
|
(predict for NP_179469.1)
|
|
Subcellular
localization
TargetP |
|
chlo 6,
other 6
|
(predict for NP_001325249.1)
|
|
chlo 6,
other 6
|
(predict for NP_179469.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
Genes directly connected with PHYB on the network
| coex z* |
Locus |
Function* |
CoexViewer |
Entrez Gene ID* |
| 5.2 |
BIM1 |
basic helix-loop-helix (bHLH) DNA-binding superfamily protein |
[detail] |
830708 |
| 4.6 |
ACC1 |
acetyl-CoA carboxylase 1 |
[detail] |
840521 |
| 4.5 |
ABCB1 |
ATP binding cassette subfamily B1 |
[detail] |
818265 |
|
Coexpressed
gene list |
[Coexpressed gene list for PHYB]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
266065_at
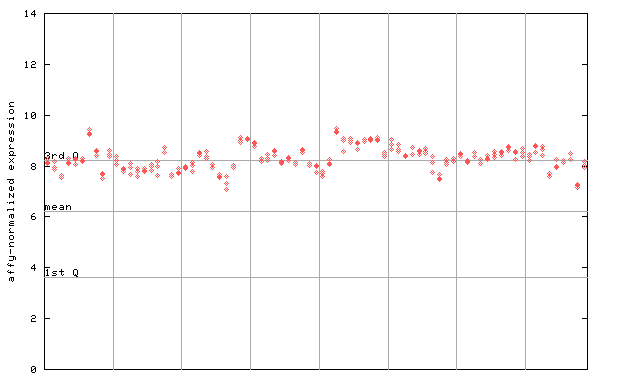
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
266065_at
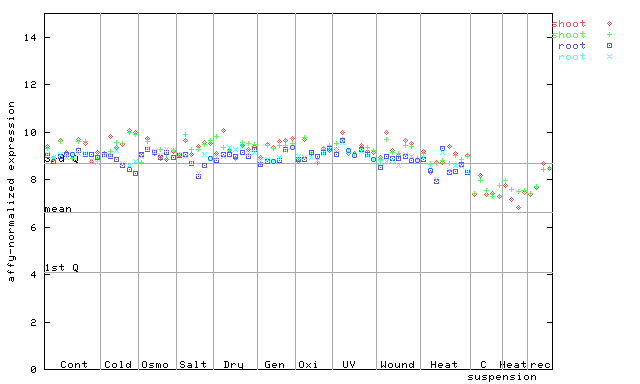
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
266065_at
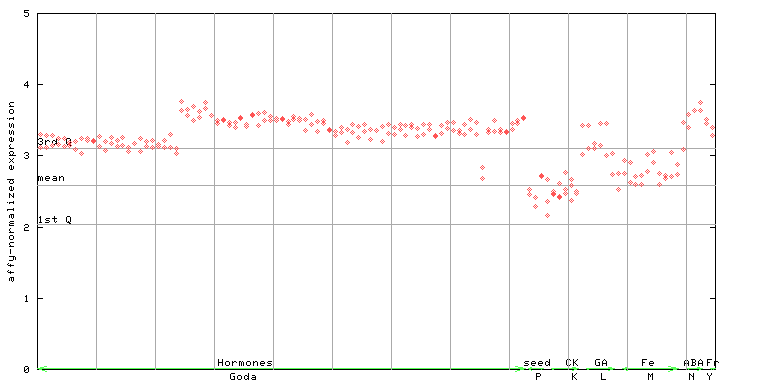
X axis is samples (xls file), and Y axis is log-expression.
|













