| functional annotation |
| Function |
COP9-signalosome 5B |


|
| GO BP |
|
GO:0010971 [list] [network] positive regulation of G2/M transition of mitotic cell cycle
|
(5 genes)
|
IMP
|
 |
|
GO:0010387 [list] [network] COP9 signalosome assembly
|
(6 genes)
|
IMP
|
 |
|
GO:2000082 [list] [network] regulation of L-ascorbic acid biosynthetic process
|
(7 genes)
|
IMP
|
 |
|
GO:0000338 [list] [network] protein deneddylation
|
(11 genes)
|
IBA
IGI
|
 |
|
GO:0010100 [list] [network] negative regulation of photomorphogenesis
|
(13 genes)
|
IGI
|
 |
|
GO:0045732 [list] [network] positive regulation of protein catabolic process
|
(45 genes)
|
IMP
|
 |
|
GO:0009585 [list] [network] red, far-red light phototransduction
|
(54 genes)
|
IEA
|
 |
|
GO:0009640 [list] [network] photomorphogenesis
|
(71 genes)
|
TAS
|
 |
|
GO:0009733 [list] [network] response to auxin
|
(407 genes)
|
IGI
|
 |
|
GO:0007275 [list] [network] multicellular organism development
|
(2714 genes)
|
IEA
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0019784 [list] [network] NEDD8-specific protease activity
|
(5 genes)
|
IBA
|
 |
|
GO:0004222 [list] [network] metalloendopeptidase activity
|
(48 genes)
|
IEA
|
 |
|
GO:0008237 [list] [network] metallopeptidase activity
|
(69 genes)
|
IBA
|
 |
|
GO:0046872 [list] [network] metal ion binding
|
(3180 genes)
|
IEA
|
 |
|
GO:0005515 [list] [network] protein binding
|
(4605 genes)
|
IPI
|
 |
|
| KEGG |
|
|
| Protein |
NP_001321294.1
NP_177279.1
|
| BLAST |
NP_001321294.1
NP_177279.1
|
| Orthologous |
[Ortholog page]
csn5 (sly)
CSN5A (ath)
LOC4337249 (osa)
LOC7458739 (ppo)
LOC11433036 (mtr)
LOC100265887 (vvi)
LOC100283276 (zma)
LOC100784909 (gma)
LOC100817371 (gma)
LOC101256005 (sly)
LOC103829303 (bra)
LOC103840816 (bra)
|
Subcellular
localization
wolf |
|
nucl 5,
cyto_nucl 4,
cyto 1,
pero 1,
plas 1,
extr 1
|
(predict for NP_001321294.1)
|
|
nucl 4,
cyto 2,
pero 2,
cysk_nucl 2,
cyto_pero 2
|
(predict for NP_177279.1)
|
|
Subcellular
localization
TargetP |
|
other 8
|
(predict for NP_001321294.1)
|
|
other 9
|
(predict for NP_177279.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath03430 |
Mismatch repair |
3 |
 |
| ath03030 |
DNA replication |
3 |
 |
| ath04120 |
Ubiquitin mediated proteolysis |
2 |
 |
| ath03420 |
Nucleotide excision repair |
2 |
 |
| ath03440 |
Homologous recombination |
2 |
 |
Genes directly connected with CSN5B on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 4.3 |
AT3G55490 |
GINS complex protein |
[detail] |
5008088 |
| 4.1 |
ZCF61 |
E3 Ubiquitin ligase family protein |
[detail] |
842247 |
| 4.0 |
AT5G06590 |
hypothetical protein |
[detail] |
830547 |
|
Coexpressed
gene list |
[Coexpressed gene list for CSN5B]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
259900_at
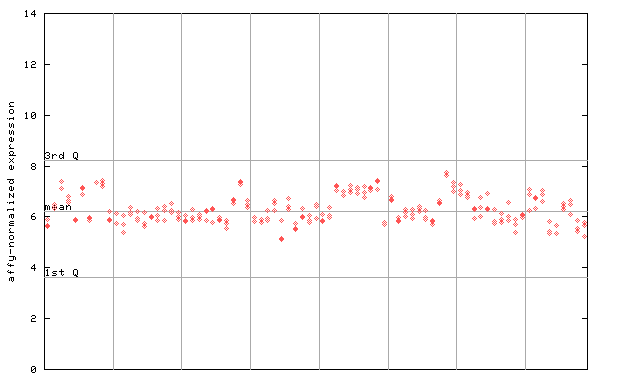
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
259900_at
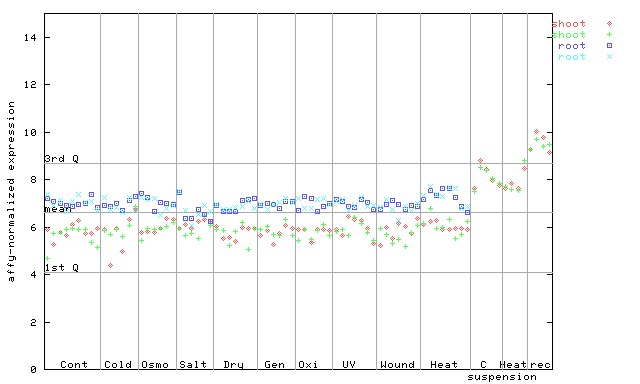
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
259900_at
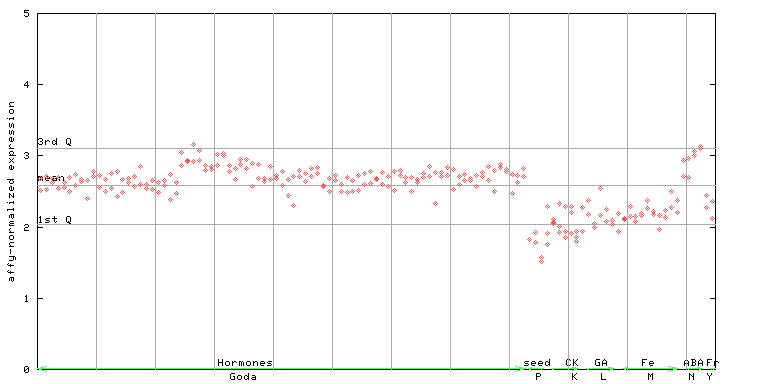
X axis is samples (xls file), and Y axis is log-expression.
|















