| functional annotation |
| Function |
gigantea protein (GI) |


|
| GO BP |
|
GO:0010378 [list] [network] temperature compensation of the circadian clock
|
(1 genes)
|
IMP
|
 |
|
GO:0048578 [list] [network] positive regulation of long-day photoperiodism, flowering
|
(9 genes)
|
IMP
|
 |
|
GO:0048586 [list] [network] regulation of long-day photoperiodism, flowering
|
(25 genes)
|
IBA
|
 |
|
GO:0042542 [list] [network] response to hydrogen peroxide
|
(52 genes)
|
IMP
|
 |
|
GO:0009585 [list] [network] red, far-red light phototransduction
|
(54 genes)
|
IEA
|
 |
|
GO:0010218 [list] [network] response to far red light
|
(54 genes)
|
IMP
|
 |
|
GO:0042752 [list] [network] regulation of circadian rhythm
|
(57 genes)
|
IBA
IMP
|
 |
|
GO:0009637 [list] [network] response to blue light
|
(98 genes)
|
IDA
|
 |
|
GO:0007623 [list] [network] circadian rhythm
|
(111 genes)
|
IDA
|
 |
|
GO:0080167 [list] [network] response to karrikin
|
(128 genes)
|
IEP
|
 |
|
GO:0009409 [list] [network] response to cold
|
(411 genes)
|
IMP
|
 |
|
GO:0009908 [list] [network] flower development
|
(447 genes)
|
TAS
|
 |
|
GO:0030154 [list] [network] cell differentiation
|
(695 genes)
|
IEA
|
 |
|
GO:0006355 [list] [network] regulation of transcription, DNA-templated
|
(1984 genes)
|
IDA
|
 |
|
| GO CC |
|
| GO MF |
|
| KEGG |
ath04712 [list] [network] Circadian rhythm - plant (36 genes) |
 |
| Protein |
NP_564180.1
|
| BLAST |
NP_564180.1
|
| Orthologous |
[Ortholog page]
LOC4325329 (osa)
LOC7492485 (ppo)
LOC11410562 (mtr)
LOC100147733 (zma)
LOC100254197 (vvi)
LOC100272803 (zma)
GIGANTEA (gma)
E2 (gma)
LOC101255788 (sly)
LOC101258346 (sly)
LOC103249166 (bra)
|
Subcellular
localization
wolf |
|
plas 5,
nucl 2,
vacu 1,
extr 1,
E.R. 1
|
(predict for NP_564180.1)
|
|
Subcellular
localization
TargetP |
|
other 6
|
(predict for NP_564180.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath04712 |
Circadian rhythm - plant |
5 |
 |
Genes directly connected with GI on the network
| coex z* |
Locus |
Function* |
Coexpression
detail |
Entrez Gene ID* |
| 12.1 |
PRR7 |
pseudo-response regulator 7 |
[detail] |
831793 |
| 8.6 |
PRR5 |
two-component response regulator-like protein |
[detail] |
832518 |
| 7.9 |
AT5G64170 |
dentin sialophosphoprotein-like protein |
[detail] |
836538 |
| 6.4 |
PDC2 |
pyruvate decarboxylase-2 |
[detail] |
835587 |
| 6.1 |
SMO2-1 |
sterol 4-alpha-methyl-oxidase 2-1 |
[detail] |
837254 |
| 5.3 |
AT3G46630 |
DCL protein (DUF3223) |
[detail] |
823816 |
| 4.5 |
AT1G21670 |
DPP6 amino-terminal domain protein |
[detail] |
838769 |
|
Coexpressed
gene list |
[Coexpressed gene list for GI]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
264211_at
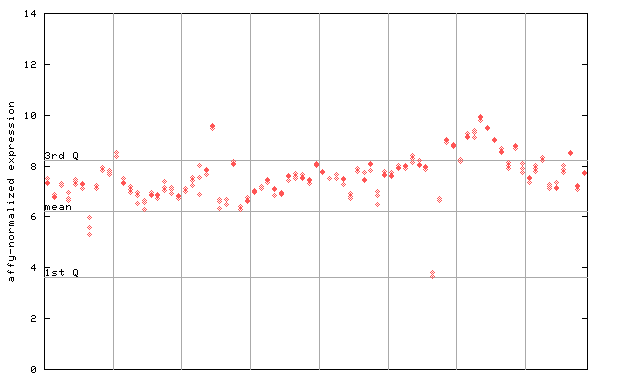
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
264211_at
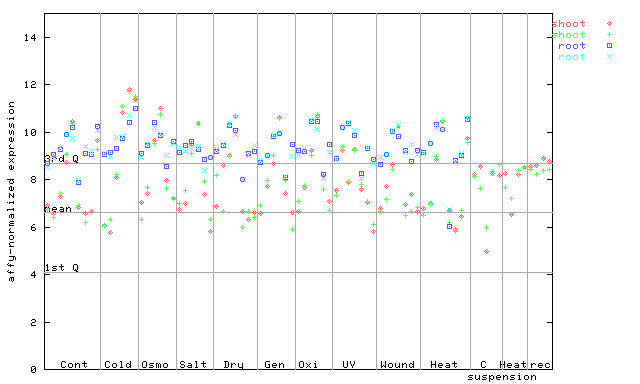
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
264211_at
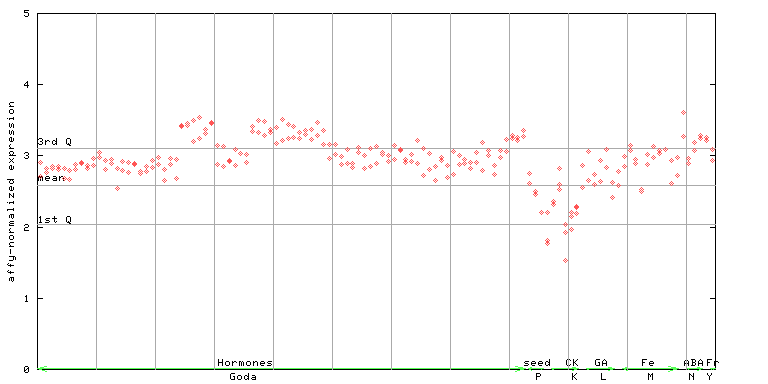
X axis is samples (xls file), and Y axis is log-expression.
|











