| functional annotation |
| Function |
Leucine-rich repeat protein kinase family protein |

 Plant GARDEN Plant GARDEN JBrowse
Plant GARDEN Plant GARDEN JBrowse
|
| GO BP |
|
GO:0006468 [list] [network] protein phosphorylation
|
(639 genes)
|
IEA
|
 |
|
GO:0016310 [list] [network] phosphorylation
|
(785 genes)
|
ISS
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0004674 [list] [network] protein serine/threonine kinase activity
|
(454 genes)
|
IEA
|
 |
|
GO:0005515 [list] [network] protein binding
|
(5066 genes)
|
IPI
|
 |
|
| KEGG |
|
|
| Protein |
NP_175593.2
|
| BLAST |
NP_175593.2
|
| Orthologous |
[Ortholog page]
AT2G04300 (ath)
AT2G14440 (ath)
AT2G14510 (ath)
FRK1 (ath)
AT2G19210 (ath)
AT2G19230 (ath)
AT2G28960 (ath)
AT2G28970 (ath)
AT2G28990 (ath)
AT2G29000 (ath)
AT3G21340 (ath)
MEE39 (ath)
AT3G46340 (ath)
AT3G46370 (ath)
AT3G46400 (ath)
AT3G46420 (ath)
AT4G20450 (ath)
RHS16 (ath)
AT4G29450 (ath)
AT4G29990 (ath)
AT5G16900 (ath)
AT5G59650 (ath)
AT5G59670 (ath)
AT5G59680 (ath)
AT1G05700 (ath)
AT1G07550 (ath)
AT1G07560 (ath)
AT1G49100 (ath)
AT1G51790 (ath)
IOS1 (ath)
AT1G51805 (ath)
AT1G51820 (ath)
AT1G51830 (ath)
AT1G51850 (ath)
AT1G51860 (ath)
AT1G51870 (ath)
RHS6 (ath)
AT1G51890 (ath)
AT1G51910 (ath)
LOC4328315 (osa)
LOC4338207 (osa)
LOC4339370 (osa)
LOC4339371 (osa)
LOC4339374 (osa)
LOC4339376 (osa)
LOC4339378 (osa)
LOC4346819 (osa)
LOC4346820 (osa)
LOC4346821 (osa)
LOC4346822 (osa)
LOC4346823 (osa)
LOC4346825 (osa)
LOC4346833 (osa)
LOC4346837 (osa)
LOC4346841 (osa)
LOC7494617 (ppo)
LOC9266163 (osa)
LOC9267814 (osa)
LOC9267979 (osa)
LOC9272354 (osa)
LOC11421236 (mtr)
LOC11422007 (mtr)
LOC11425516 (mtr)
LOC11429446 (mtr)
LOC11433438 (mtr)
LOC11434037 (mtr)
LOC11437028 (mtr)
LOC18108501 (ppo)
LOC18108503 (ppo)
LOC18108531 (ppo)
LOC18110286 (ppo)
LOC18110288 (ppo)
LOC18110448 (ppo)
LOC18110542 (ppo)
LOC18110545 (ppo)
LOC18110550 (ppo)
LOC25497008 (mtr)
LOC25500355 (mtr)
LOC25500358 (mtr)
LOC25500359 (mtr)
LOC25500361 (mtr)
LOC25500364 (mtr)
LOC25500366 (mtr)
LOC25500367 (mtr)
LOC25500376 (mtr)
LOC25500764 (mtr)
LOC25502465 (mtr)
SIK2 (gma)
LOC100775741 (gma)
LOC100776284 (gma)
LOC100776818 (gma)
LOC100778537 (gma)
LOC100779102 (gma)
LOC100779474 (gma)
LOC100779617 (gma)
LOC100781084 (gma)
LOC100781785 (gma)
LOC100785345 (gma)
LOC100791717 (gma)
LOC100792239 (gma)
LOC100793137 (gma)
LOC100794504 (gma)
LOC100795183 (gma)
LOC100818475 (gma)
LOC101257947 (sly)
LOC101262429 (sly)
LOC101266705 (sly)
LOC102662730 (gma)
LOC102663143 (gma)
LOC103829493 (bra)
LOC103829497 (bra)
LOC103836435 (bra)
LOC103843445 (bra)
LOC103844100 (bra)
LOC103845385 (bra)
LOC103850089 (bra)
LOC103851563 (bra)
LOC103851564 (bra)
LOC103851565 (bra)
LOC103851763 (bra)
LOC103853036 (bra)
LOC103853548 (bra)
LOC103856616 (bra)
LOC103859903 (bra)
LOC103861877 (bra)
LOC103864933 (bra)
LOC103864934 (bra)
LOC103868533 (bra)
LOC103868536 (bra)
LOC103868540 (bra)
LOC103868542 (bra)
LOC103868543 (bra)
LOC103868545 (bra)
LOC103868632 (bra)
LOC103869180 (bra)
LOC103871250 (bra)
LOC103871251 (bra)
LOC103871253 (bra)
LOC103871255 (bra)
LOC103873634 (bra)
LOC106794186 (gma)
LOC106799536 (gma)
LOC107275782 (osa)
LOC107276230 (osa)
LOC107276752 (osa)
LOC107278492 (osa)
LOC107278845 (osa)
LOC112325225 (ppo)
LOC112416101 (mtr)
LOC112417465 (mtr)
LOC112936039 (osa)
LOC117125683 (bra)
LOC120575758 (mtr)
LOC123043522 (tae)
LOC123044006 (tae)
LOC123044087 (tae)
LOC123044243 (tae)
LOC123054720 (tae)
LOC123063692 (tae)
LOC123067573 (tae)
LOC123079142 (tae)
LOC123080188 (tae)
LOC123080196 (tae)
LOC123080205 (tae)
LOC123080238 (tae)
LOC123083057 (tae)
LOC123084992 (tae)
LOC123085003 (tae)
LOC123086249 (tae)
LOC123094168 (tae)
LOC123094169 (tae)
LOC123096526 (tae)
LOC123096537 (tae)
LOC123096545 (tae)
LOC123096554 (tae)
LOC123096599 (tae)
LOC123096617 (tae)
LOC123096813 (tae)
LOC123097489 (tae)
LOC123103544 (tae)
LOC123106972 (tae)
LOC123114350 (tae)
LOC123114773 (tae)
LOC123116086 (tae)
LOC123120524 (tae)
LOC123124720 (tae)
LOC123126207 (tae)
LOC123132270 (tae)
LOC123132271 (tae)
LOC123132273 (tae)
LOC123136316 (tae)
LOC123136318 (tae)
LOC123136319 (tae)
LOC123138953 (tae)
LOC123140966 (tae)
LOC123143830 (tae)
LOC123143832 (tae)
LOC123143833 (tae)
LOC123143893 (tae)
LOC123157067 (tae)
LOC123160036 (tae)
LOC123160179 (tae)
LOC123160205 (tae)
LOC123160231 (tae)
LOC123162979 (tae)
LOC123162990 (tae)
LOC123173379 (tae)
LOC123173395 (tae)
LOC123173449 (tae)
LOC123179719 (tae)
LOC123179738 (tae)
LOC123179780 (tae)
LOC123180475 (tae)
LOC123182688 (tae)
LOC123182689 (tae)
LOC123395140 (hvu)
LOC123398290 (hvu)
LOC123401280 (hvu)
LOC123401354 (hvu)
LOC123401635 (hvu)
LOC123404089 (hvu)
LOC123404264 (hvu)
LOC123404421 (hvu)
LOC123404434 (hvu)
LOC123404717 (hvu)
LOC123408210 (hvu)
LOC123416831 (hvu)
LOC123416847 (hvu)
LOC123416895 (hvu)
LOC123440187 (hvu)
LOC123442690 (hvu)
LOC123448982 (hvu)
LOC123450805 (hvu)
LOC123450806 (hvu)
|
Subcellular
localization
wolf |
|
chlo 3,
E.R. 2,
nucl 1,
cyto 1,
cyto_nucl 1,
E.R._vacu 1,
E.R._plas 1
|
(predict for NP_175593.2)
|
|
Subcellular
localization
TargetP |
|
other 8
|
(predict for NP_175593.2)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
|
Coexpressed
gene list |
[Coexpressed gene list for AT1G51810]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
256180_at
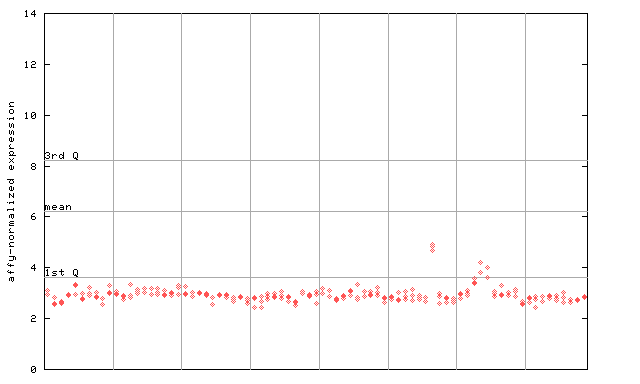
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
256180_at
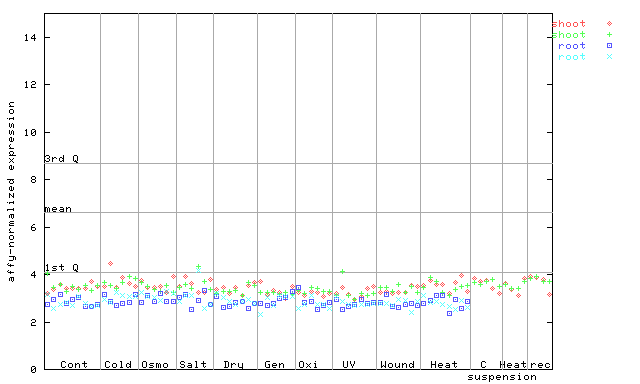
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
256180_at
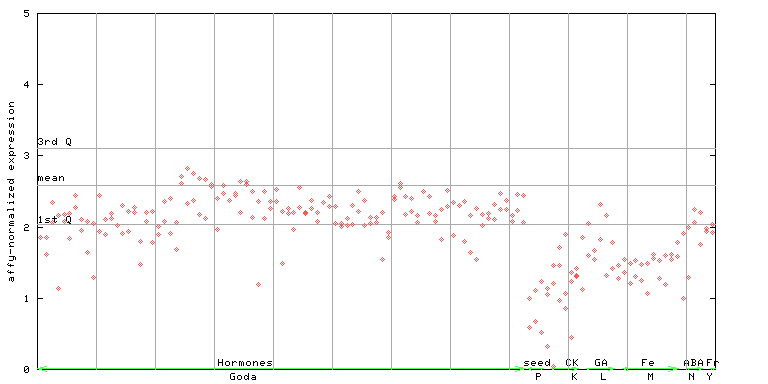
X axis is samples (xls file), and Y axis is log-expression.
|





