| functional annotation |
| Function |
Peroxidase superfamily protein |




|
| GO BP |
|
GO:0001561 [list] [network] fatty acid alpha-oxidation
|
(2 genes)
|
IDA
|
 |
|
GO:1902609 [list] [network] (R)-2-hydroxy-alpha-linolenic acid biosynthetic process
|
(2 genes)
|
IMP
|
 |
|
GO:0071732 [list] [network] cellular response to nitric oxide
|
(4 genes)
|
IEP
|
 |
|
GO:0034614 [list] [network] cellular response to reactive oxygen species
|
(29 genes)
|
IEP
|
 |
|
GO:0071446 [list] [network] cellular response to salicylic acid stimulus
|
(110 genes)
|
IEP
|
 |
|
GO:0009627 [list] [network] systemic acquired resistance
|
(117 genes)
|
IMP
|
 |
|
GO:0008219 [list] [network] cell death
|
(126 genes)
|
IEP
|
 |
|
GO:0009751 [list] [network] response to salicylic acid
|
(438 genes)
|
IEP
|
 |
|
GO:0006979 [list] [network] response to oxidative stress
|
(545 genes)
|
IMP
|
 |
|
GO:0050832 [list] [network] defense response to fungus
|
(705 genes)
|
IDA
|
 |
|
GO:0042742 [list] [network] defense response to bacterium
|
(980 genes)
|
IEP
|
 |
|
GO:0009737 [list] [network] response to abscisic acid
|
(1086 genes)
|
IMP
|
 |
|
GO:0006629 [list] [network] lipid metabolic process
|
(1087 genes)
|
TAS
|
 |
|
| GO CC |
|
GO:0012511 [list] [network] monolayer-surrounded lipid storage body
|
(9 genes)
|
IDA
|
 |
|
GO:0005576 [list] [network] extracellular region
|
(3154 genes)
|
ISM
|
 |
|
| GO MF |
|
GO:0016702 [list] [network] oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen
|
(29 genes)
|
IDA
|
 |
|
| KEGG |
ath00592 [list] [network] alpha-Linolenic acid metabolism (43 genes) |
 |
| Protein |
NP_186791.1
|
| BLAST |
NP_186791.1
|
| Orthologous |
[Ortholog page]
LOC543895 (sly)
LOC543896 (sly)
LOC4352160 (osa)
LOC7485908 (ppo)
LOC25495214 (mtr)
LOC25495215 (mtr)
LOC25495216 (mtr)
LOC100794600 (gma)
LOC100797235 (gma)
LOC100817200 (gma)
LOC101246396 (sly)
LOC101290585 (tae)
LOC103870978 (bra)
LOC123072527 (tae)
LOC123072528 (tae)
LOC123408826 (hvu)
LOC123412509 (hvu)
|
Subcellular
localization
wolf |
|
cyto 5,
chlo 2,
extr 1,
vacu 1,
pero 1,
mito_plas 1
|
(predict for NP_186791.1)
|
|
Subcellular
localization
TargetP |
|
scret 9
|
(predict for NP_186791.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
Genes directly connected with DOX1 on the network
| coex z* |
Locus |
Function* |
CoexViewer |
Entrez Gene ID* |
| 14.0 |
LAC7 |
laccase 7 |
[detail] |
820078 |
| 11.2 |
OSM34 |
osmotin 34 |
[detail] |
826770 |
| 10.4 |
KTI1 |
kunitz trypsin inhibitor 1 |
[detail] |
843660 |
| 9.7 |
AT1G30700 |
FAD-binding Berberine family protein |
[detail] |
839950 |
| 8.4 |
UPI |
Serine protease inhibitor, potato inhibitor I-type family protein |
[detail] |
834378 |
| 8.3 |
PDE337 |
VQ motif-containing protein |
[detail] |
843174 |
| 6.3 |
PLP1 |
Acyl transferase/acyl hydrolase/lysophospholipase superfamily protein |
[detail] |
829861 |
| 5.0 |
LOX1 |
lipoxygenase 1 |
[detail] |
841944 |
| 5.0 |
ARSK1 |
root-specific kinase 1 |
[detail] |
817169 |
| 4.3 |
PAP8 |
purple acid phosphatase 8 |
[detail] |
814720 |
|
Coexpressed
gene list |
[Coexpressed gene list for DOX1]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
258957_at
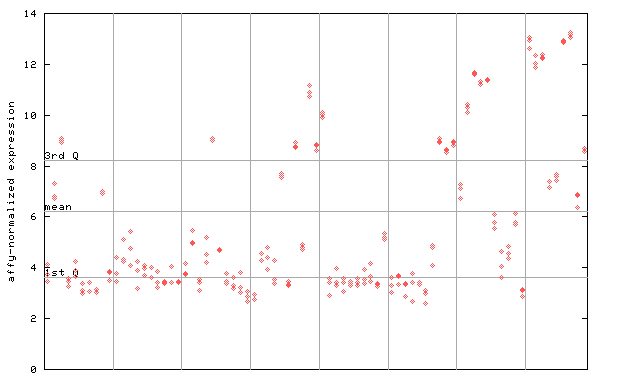
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
258957_at
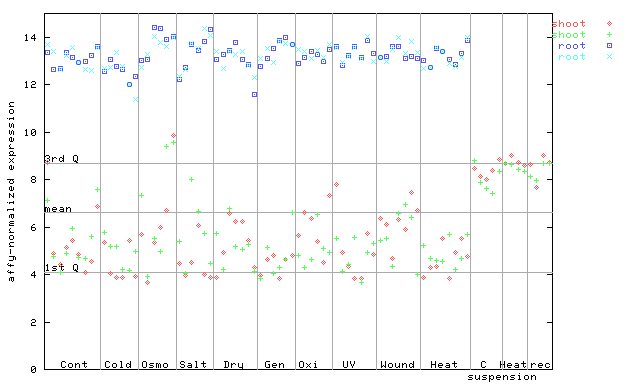
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
258957_at
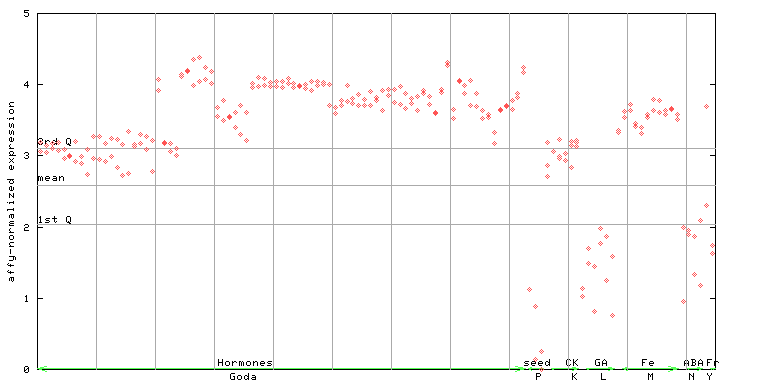
X axis is samples (xls file), and Y axis is log-expression.
|












