| functional annotation |
| Function |
Homeodomain-like superfamily protein |




|
| GO BP |
|
GO:0032922 [list] [network] circadian regulation of gene expression
|
(9 genes)
|
IMP
|
 |
|
GO:0045944 [list] [network] positive regulation of transcription by RNA polymerase II
|
(11 genes)
|
IMP
|
 |
|
GO:0042753 [list] [network] positive regulation of circadian rhythm
|
(12 genes)
|
IMP
|
 |
|
GO:0043966 [list] [network] histone H3 acetylation
|
(17 genes)
|
IMP
|
 |
|
GO:0042752 [list] [network] regulation of circadian rhythm
|
(61 genes)
|
IGI
|
 |
|
GO:0010628 [list] [network] positive regulation of gene expression
|
(71 genes)
|
IMP
|
 |
|
GO:0048573 [list] [network] photoperiodism, flowering
|
(94 genes)
|
IMP
|
 |
|
GO:0006355 [list] [network] regulation of DNA-templated transcription
|
(1656 genes)
|
TAS
|
 |
|
| GO CC |
|
| GO MF |
|
GO:0043565 [list] [network] sequence-specific DNA binding
|
(890 genes)
|
IDA
|
 |
|
GO:0003700 [list] [network] DNA-binding transcription factor activity
|
(1576 genes)
|
ISS
|
 |
|
| KEGG |
|
|
| Protein |
NP_001030662.1
NP_001327345.1
NP_001327346.1
NP_001327347.1
NP_001327348.1
NP_001327349.1
NP_001327350.1
NP_001327351.1
NP_187571.2
|
| BLAST |
NP_001030662.1
NP_001327345.1
NP_001327346.1
NP_001327347.1
NP_001327348.1
NP_001327349.1
NP_001327350.1
NP_001327351.1
NP_187571.2
|
| Orthologous |
[Ortholog page]
MYB118 (gma)
LCL1 (ath)
LOC7467599 (ppo)
LOC18100248 (ppo)
LOC25484055 (mtr)
LOC100783482 (gma)
LOC100797466 (gma)
LOC100807156 (gma)
LOC101253545 (sly)
LOC103847419 (bra)
LOC103848060 (bra)
LOC103870531 (bra)
|
Subcellular
localization
wolf |
|
nucl 9,
plas 1,
cysk 1,
cysk_plas 1
|
(predict for NP_001030662.1)
|
|
nucl 8,
chlo 1,
cyto 1,
plas 1,
cyto_plas 1
|
(predict for NP_001327345.1)
|
|
nucl 9,
chlo 1,
plas 1
|
(predict for NP_001327346.1)
|
|
nucl 9,
chlo 1,
plas 1
|
(predict for NP_001327347.1)
|
|
nucl 9,
cyto 1,
plas 1,
cyto_plas 1
|
(predict for NP_001327348.1)
|
|
nucl 9,
plas 1,
cysk 1,
cysk_plas 1
|
(predict for NP_001327349.1)
|
|
nucl 8,
chlo 1,
cyto 1,
plas 1,
cyto_plas 1
|
(predict for NP_001327350.1)
|
|
nucl 8,
cyto 1,
plas 1,
cysk 1,
cysk_plas 1,
cyto_plas 1
|
(predict for NP_001327351.1)
|
|
nucl 9,
cyto 1,
plas 1,
cyto_plas 1
|
(predict for NP_187571.2)
|
|
Subcellular
localization
TargetP |
|
chlo 7,
other 3
|
(predict for NP_001030662.1)
|
|
other 7
|
(predict for NP_001327345.1)
|
|
other 7
|
(predict for NP_001327346.1)
|
|
other 7
|
(predict for NP_001327347.1)
|
|
chlo 7,
other 3
|
(predict for NP_001327348.1)
|
|
chlo 7,
other 3
|
(predict for NP_001327349.1)
|
|
other 7
|
(predict for NP_001327350.1)
|
|
chlo 7,
other 3
|
(predict for NP_001327351.1)
|
|
chlo 7,
other 3
|
(predict for NP_187571.2)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath04712 |
Circadian rhythm - plant |
2 |
 |
Genes directly connected with RVE8 on the network
| coex z* |
Locus |
Function* |
CoexViewer |
Entrez Gene ID* |
| 16.5 |
LHY |
Homeodomain-like superfamily protein |
[detail] |
839341 |
| 13.0 |
CCA1 |
circadian clock associated 1 |
[detail] |
819296 |
| 10.6 |
AT3G54500 |
agglutinin-like protein |
[detail] |
824615 |
| 10.5 |
BBX19 |
B-box type zinc finger family protein |
[detail] |
830051 |
| 7.8 |
LCL1 |
LHY/CCA1-like 1 |
[detail] |
831766 |
| 5.8 |
AT1G70250 |
receptor serine/threonine kinase |
[detail] |
843361 |
| 4.2 |
AT2G27360 |
GDSL-like Lipase/Acylhydrolase superfamily protein |
[detail] |
817279 |
|
Coexpressed
gene list |
[Coexpressed gene list for RVE8]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
258724_at
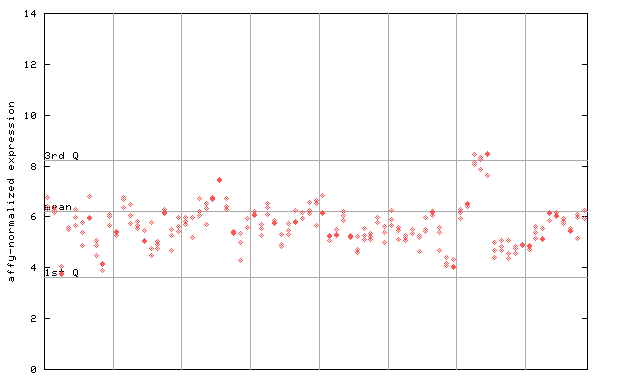
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
258724_at
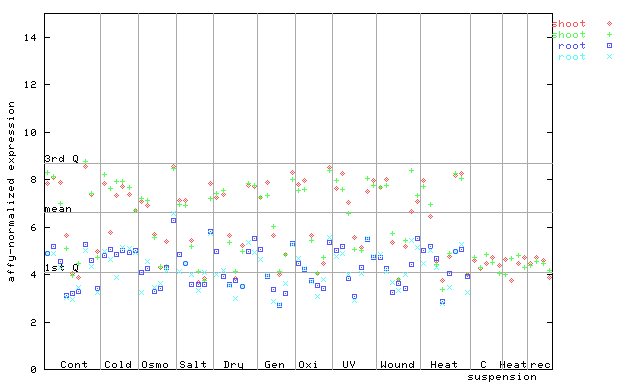
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
258724_at
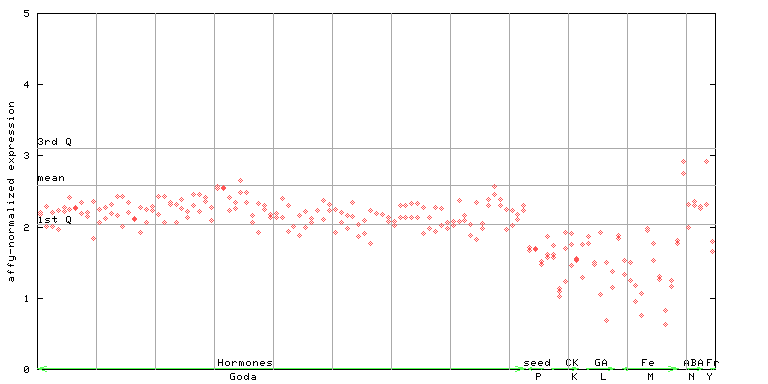
X axis is samples (xls file), and Y axis is log-expression.
|




















