| functional annotation |
| Function |
LSD1 zinc finger family protein |




|
| GO BP |
|
GO:0010602 [list] [network] regulation of 1-aminocyclopropane-1-carboxylate metabolic process
|
(1 genes)
|
IMP
|
 |
|
GO:0010618 [list] [network] aerenchyma formation
|
(3 genes)
|
IMP
|
 |
|
GO:0002240 [list] [network] response to molecule of oomycetes origin
|
(6 genes)
|
IMP
|
 |
|
GO:0000303 [list] [network] response to superoxide
|
(9 genes)
|
IMP
|
 |
|
GO:0010310 [list] [network] regulation of hydrogen peroxide metabolic process
|
(15 genes)
|
IMP
|
 |
|
GO:0009862 [list] [network] systemic acquired resistance, salicylic acid mediated signaling pathway
|
(16 genes)
|
IEP
|
 |
|
GO:0043069 [list] [network] negative regulation of programmed cell death
|
(28 genes)
|
IGI
|
 |
|
GO:0010104 [list] [network] regulation of ethylene-activated signaling pathway
|
(30 genes)
|
IMP
|
 |
|
GO:0009626 [list] [network] plant-type hypersensitive response
|
(46 genes)
|
IMP
|
 |
|
GO:0043067 [list] [network] regulation of programmed cell death
|
(67 genes)
|
IGI
|
 |
|
GO:0008219 [list] [network] cell death
|
(126 genes)
|
IMP
|
 |
|
GO:0001666 [list] [network] response to hypoxia
|
(325 genes)
|
IMP
|
 |
|
GO:0009409 [list] [network] response to cold
|
(472 genes)
|
IMP
|
 |
|
| GO CC |
|
GO:0005576 [list] [network] extracellular region
|
(3154 genes)
|
ISM
|
 |
|
| GO MF |
|
GO:0003700 [list] [network] DNA-binding transcription factor activity
|
(1576 genes)
|
TAS
|
 |
|
GO:0005515 [list] [network] protein binding
|
(5066 genes)
|
IPI
|
 |
|
| KEGG |
|
|
| Protein |
NP_001031678.1
NP_001031679.1
NP_001031680.2
NP_001078413.1
NP_001154257.1
NP_001328264.1
NP_567599.3
NP_849548.1
NP_849549.1
|
| BLAST |
NP_001031678.1
NP_001031679.1
NP_001031680.2
NP_001078413.1
NP_001154257.1
NP_001328264.1
NP_567599.3
NP_849548.1
NP_849549.1
|
| Orthologous |
[Ortholog page]
LOC542922 (tae)
LOC4333536 (osa)
LOC4352767 (osa)
LOC7455048 (ppo)
LOC7490691 (ppo)
LOC11427425 (mtr)
LOC25486678 (mtr)
LOC100527656 (gma)
LOC100776394 (gma)
LOC100776644 (gma)
LOC100792570 (gma)
LOC101253843 (sly)
LOC101265799 (sly)
LOC103834161 (bra)
LOC103858192 (bra)
LOC103861197 (bra)
LOC123102232 (tae)
LOC123119416 (tae)
LOC123122737 (tae)
LOC123395547 (hvu)
LOC123452126 (hvu)
|
Subcellular
localization
wolf |
|
chlo 9,
nucl 1,
vacu 1
|
(predict for NP_001031678.1)
|
|
chlo 9,
nucl 1,
vacu 1
|
(predict for NP_001031679.1)
|
|
chlo 9,
nucl 1,
vacu 1
|
(predict for NP_001031680.2)
|
|
chlo 9
|
(predict for NP_001078413.1)
|
|
chlo 9
|
(predict for NP_001154257.1)
|
|
chlo 7,
cyto 1,
mito 1,
cysk 1,
cyto_nucl 1,
cyto_pero 1,
cyto_E.R. 1,
cyto_plas 1
|
(predict for NP_001328264.1)
|
|
chlo 9,
nucl 1,
vacu 1
|
(predict for NP_567599.3)
|
|
chlo 9,
nucl 1,
vacu 1
|
(predict for NP_849548.1)
|
|
chlo 9
|
(predict for NP_849549.1)
|
|
Subcellular
localization
TargetP |
|
mito 2
|
(predict for NP_001031678.1)
|
|
mito 2
|
(predict for NP_001031679.1)
|
|
mito 2
|
(predict for NP_001031680.2)
|
|
other 5
|
(predict for NP_001078413.1)
|
|
mito 2
|
(predict for NP_001154257.1)
|
|
mito 2
|
(predict for NP_001328264.1)
|
|
mito 2
|
(predict for NP_567599.3)
|
|
mito 2
|
(predict for NP_849548.1)
|
|
other 5
|
(predict for NP_849549.1)
|
|
| Gene coexpression |
Network*for
coexpressed
genes |
| KEGG* ID |
Title |
#genes |
Link to the KEGG* map
(Multiple genes) |
| ath04130 |
SNARE interactions in vesicular transport |
2 |
 |
Genes directly connected with LSD1 on the network
| coex z* |
Locus |
Function* |
CoexViewer |
Entrez Gene ID* |
| 8.8 |
RPL18AA |
60S ribosomal protein L18A-1 |
[detail] |
839876 |
| 8.8 |
AT4G26400 |
RING/U-box superfamily protein |
[detail] |
828746 |
| 7.9 |
AT5G05100 |
Single-stranded nucleic acid binding R3H protein |
[detail] |
830392 |
| 7.0 |
AT1G70590 |
F-box family protein |
[detail] |
843396 |
| 6.8 |
AT2G21120 |
magnesium transporter, putative (DUF803) |
[detail] |
816647 |
| 6.3 |
AT4G24730 |
manganese-dependent ADP-ribose/CDP-alcohol diphosphatase-like protein |
[detail] |
828575 |
| 5.2 |
AT3G06760 |
Drought-responsive family protein |
[detail] |
819861 |
|
Coexpressed
gene list |
[Coexpressed gene list for LSD1]
|
| Gene expression |
| All samples |
[Expression pattern for all samples]
|
AtGenExpress*
(Development) |
254477_at
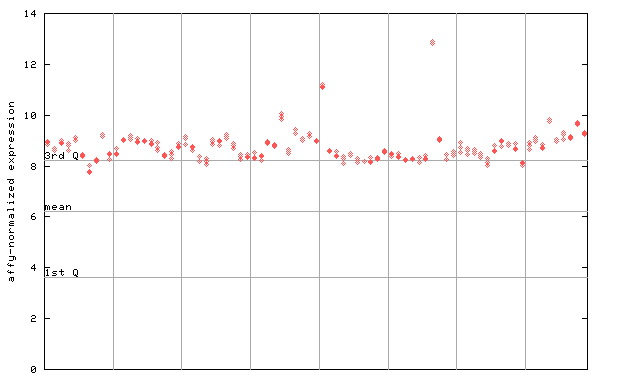
X axis is samples (pdf file), and Y axis is log2-expression.
"1st Q", "mean" and "3rd Q" indecate the values for all genes on a GeneChip. Q: quartile.
|
AtGenExpress*
(Stress) |
254477_at
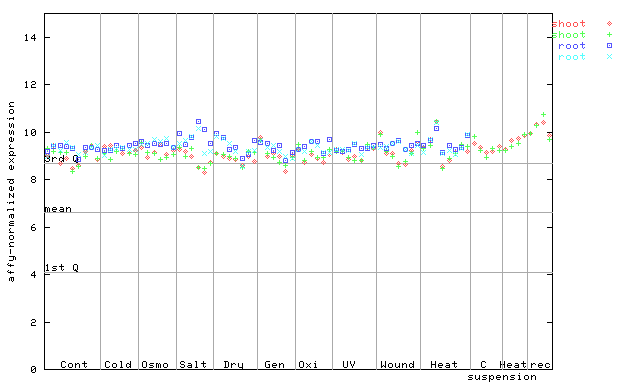
X axis is samples (pdf file), and Y axis is log2-expression.
|
AtGenExpress*
(Hormone) |
254477_at
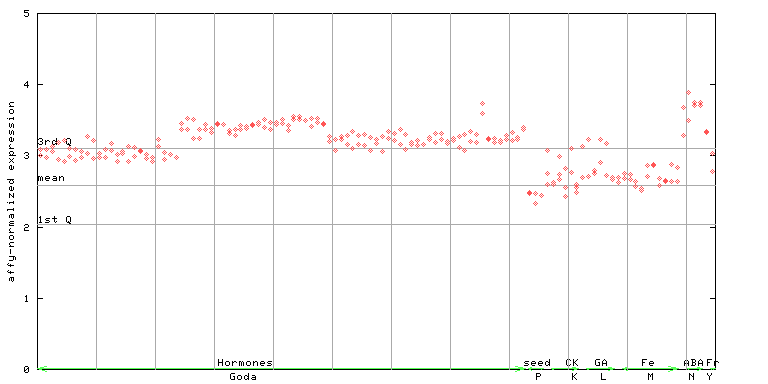
X axis is samples (xls file), and Y axis is log-expression.
|




















