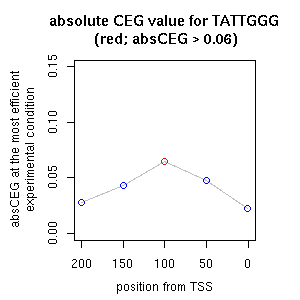
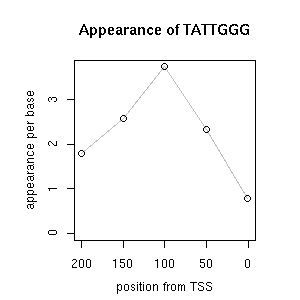
| reported seq | reported name | reference |
|---|---|---|
| no correspondence | ||
 |
 |
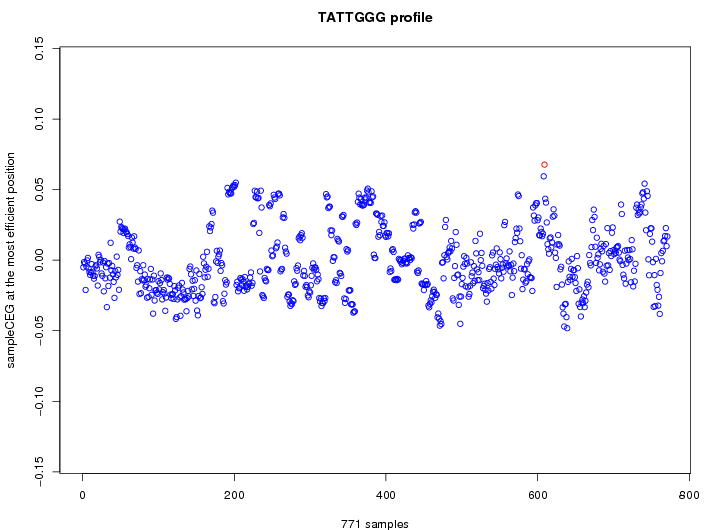
| GO ID | term | -Log(p) | number of {A; genes in a GO} | numver of {B; genes with this heptamer} | {A} x {B} |
|---|---|---|---|---|---|
| no functional bias detected | |||||
| correl | TF | function ((alias))* | TF family (link to AGRIS) |
|---|---|---|---|
| 0.58 | At4g30930 | 50S ribosomal protein L21, mitochondrial (RPL21M) ((WRKY32)) | WRKY |
| 0.58 | At1g71260 | expressed protein | WHIRLY |
| 0.53 | At3g52910 | expressed protein | GRF |
| 0.50 | At3g57150 | dyskerin, putative / nucleolar protein NAP57, putative ((NAP57)) | NAC |
| 0.49 | At5g16070 | chaperonin, putative | TCP |
| 0.48 | At1g14410 | DNA-binding protein-related | WHIRLY |
| -0.45 | At3g19580 | zinc finger (C2H2 type) protein 2 (AZF2) ((AZF2)) | C2H2 |
| -0.44 | At3g52800 | zinc finger (AN1-like) family protein | C2H2 |
| 0.44 | At5g05610 | PHD finger family protein | Alfin-like |
| -0.44 | At5g66070 | zinc finger (C3HC4-type RING finger) family protein | C3H |
| 0.44 | At5g02470 | DP-2 transcription factor, putative (DPA) ((DPA)) | E2F-DP |
| -0.43 | At5g55970 | zinc finger (C3HC4-type RING finger) family protein | C3H |
| 0.43 | At2g02740 | transcription factor, putative | WHIRLY |
| 0.43 | At3g16870 | zinc finger (GATA type) family protein | C2C2-Gata |
| 0.43 | At3g06740 | zinc finger (GATA type) family protein | C2C2-Gata |
|